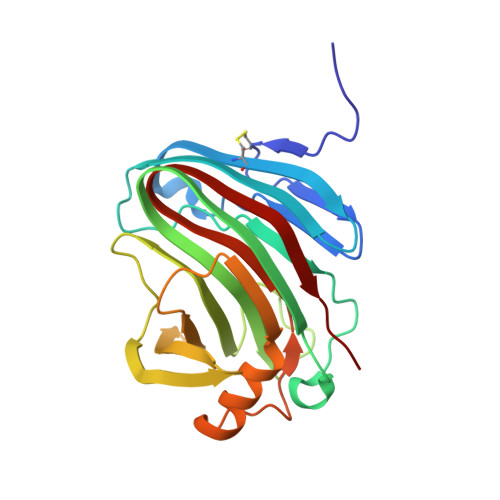
X-ray Structure and Molecular Dynamics Simulations of Endoglucanase 3 from Trichoderma harzianum: Structural Organization and Substrate Recognition by Endoglucanases That Lack Cellulose Binding Module.
Prates, E.T., Stankovic, I., Silveira, R.L., Liberato, M.V., Henrique-Silva, F., Pereira, N., Polikarpov, I., Skaf, M.S.(2013) PLoS One 8: e59069-e59069
- PubMed: 23516599 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0059069
- Primary Citation Related Structures:
4H7M - PubMed Abstract:
Plant biomass holds a promise for the production of second-generation ethanol via enzymatic hydrolysis, but its utilization as a biofuel resource is currently limited to a large extent by the cost and low efficiency of the cellulolytic enzymes. Considerable efforts have been dedicated to elucidate the mechanisms of the enzymatic process. It is well known that most cellulases possess a catalytic core domain and a carbohydrate binding module (CBM), without which the enzymatic activity can be drastically reduced. However, Cel12A members of the glycosyl hydrolases family 12 (GHF12) do not bear a CBM and yet are able to hydrolyze amorphous cellulose quite efficiently. Here, we use X-ray crystallography and molecular dynamics simulations to unravel the molecular basis underlying the catalytic capability of endoglucanase 3 from Trichoderma harzianum (ThEG3), a member of the GHF12 enzymes that lacks a CBM. A comparative analysis with the Cellulomonas fimi CBM identifies important residues mediating interactions of EG3s with amorphous regions of the cellulose. For instance, three aromatic residues constitute a harboring wall of hydrophobic contacts with the substrate in both ThEG3 and CfCBM structures. Moreover, residues at the entrance of the active site cleft of ThEG3 are identified, which might hydrogen bond to the substrate. We advocate that the ThEG3 residues Asn152 and Glu201 interact with the substrate similarly to the corresponding CfCBM residues Asn81 and Arg75. Altogether, these results show that CBM motifs are incorporated within the ThEG3 catalytic domain and suggest that the enzymatic efficiency is associated with the length and position of the substrate chain, being higher when the substrate interact with the aromatic residues at the entrance of the cleft and the catalytic triad. Our results provide guidelines for rational protein engineering aiming to improve interactions of GHF12 enzymes with cellulosic substrates.
- Institute of Chemistry, State University of Campinas-UNICAMP. Cx.P. 6154, Campinas, São Paulo, Brazil.
Organizational Affiliation: