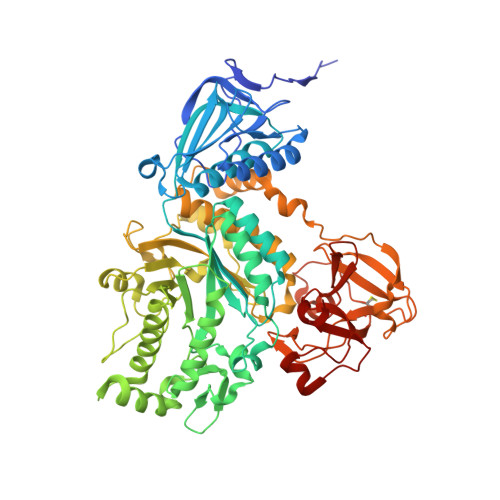
Crystal structures of a glycoside hydrolase family 20 lacto-N-biosidase from Bifidobacterium bifidum
Ito, T., Katayama, T., Hattie, M., Sakurama, H., Wada, J., Suzuki, R., Ashida, H., Wakagi, T., Yamamoto, K., Stubbs, K.A., Fushinobu, S.(2013) J Biological Chem 288: 11795-11806
- PubMed: 23479733 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M112.420109
- Primary Citation Related Structures:
4H04, 4JAW - PubMed Abstract:
Human milk oligosaccharides contain a large variety of oligosaccharides, of which lacto-N-biose I (Gal-β1,3-GlcNAc; LNB) predominates as a major core structure. A unique metabolic pathway specific for LNB has recently been identified in the human commensal bifidobacteria. Several strains of infant gut-associated bifidobacteria possess lacto-N-biosidase, a membrane-anchored extracellular enzyme, that liberates LNB from the nonreducing end of human milk oligosaccharides and plays a key role in the metabolic pathway of these compounds. Lacto-N-biosidase belongs to the glycoside hydrolase family 20, and its reaction proceeds via a substrate-assisted catalytic mechanism. Several crystal structures of GH20 β-N-acetylhexosaminidases, which release monosaccharide GlcNAc from its substrate, have been determined, but to date, a structure of lacto-N-biosidase is unknown. Here, we have determined the first three-dimensional structures of lacto-N-biosidase from Bifidobacterium bifidum JCM1254 in complex with LNB and LNB-thiazoline (Gal-β1,3-GlcNAc-thiazoline) at 1.8-Å resolution. Lacto-N-biosidase consists of three domains, and the C-terminal domain has a unique β-trefoil-like fold. Compared with other β-N-acetylhexosaminidases, lacto-N-biosidase has a wide substrate-binding pocket with a -2 subsite specific for β-1,3-linked Gal, and the residues responsible for Gal recognition were identified. The bound ligands are recognized by extensive hydrogen bonds at all of their hydroxyls consistent with the enzyme's strict substrate specificity for the LNB moiety. The GlcNAc sugar ring of LNB is in a distorted conformation near (4)E, whereas that of LNB-thiazoline is in a (4)C1 conformation. A possible conformational pathway for the lacto-N-biosidase reaction is discussed.
- Department of Biotechnology, University of Tokyo, 1-1-1 Yayoi, Bunkyo-ku, Tokyo 113-8657, Japan.
Organizational Affiliation: