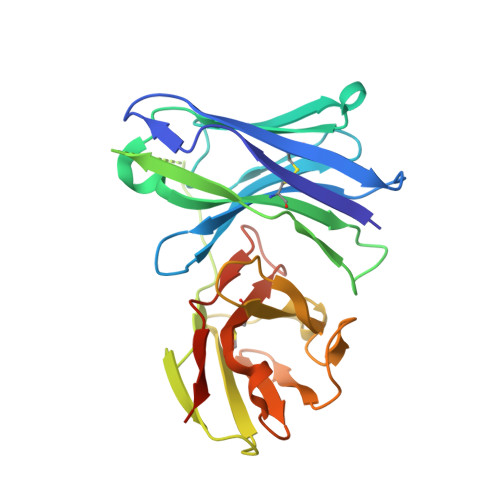
Affinity improvement of a therapeutic antibody to methamphetamine and amphetamine through structure-based antibody engineering.
Thakkar, S., Nanaware-Kharade, N., Celikel, R., Peterson, E.C., Varughese, K.I.(2014) Sci Rep 4: 3673-3673
- PubMed: 24419156 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/srep03673
- Primary Citation Related Structures:
4GQP - PubMed Abstract:
Methamphetamine (METH) abuse is a worldwide threat, without any FDA approved medications. Anti-METH IgGs and single chain fragments (scFvs) have shown efficacy in preclinical studies. Here we report affinity enhancement of an anti-METH scFv for METH and its active metabolite amphetamine (AMP), through the introduction of point mutations, rationally designed to optimize the shape and hydrophobicity of the antibody binding pocket. The binding affinity was measured using saturation binding technique. The mutant scFv-S93T showed 3.1 fold enhancement in affinity for METH and 26 fold for AMP. The scFv-I37M and scFv-Y34M mutants showed enhancement of 94, and 8 fold for AMP, respectively. Structural analysis of scFv-S93T:METH revealed that the substitution of Ser residue by Thr caused the expulsion of a water molecule from the cavity, creating a more hydrophobic environment for the binding that dramatically increases the affinities for METH and AMP.
- Department of Physiology and Biophysics, College of Medicine, University of Arkansas for Medical Sciences, Little Rock, Arkansas, USA.
Organizational Affiliation: