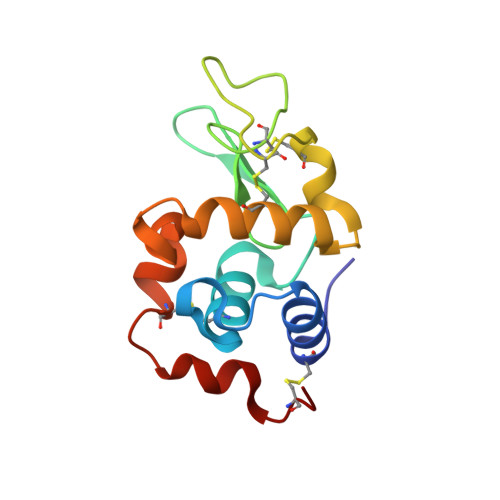
Room-temperature X-ray diffraction studies of cisplatin and carboplatin binding to His15 of HEWL after prolonged chemical exposure.
Tanley, S.W., Schreurs, A.M., Kroon-Batenburg, L.M., Helliwell, J.R.(2012) Acta Crystallogr Sect F Struct Biol Cryst Commun 68: 1300-1306
- PubMed: 23143236 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S1744309112042005
- Primary Citation Related Structures:
4G49, 4G4B, 4G4C, 4G4H - PubMed Abstract:
The anticancer complexes cisplatin and carboplatin are known to bind to both the Nδ and the Nℇ atoms of His15 of hen egg-white lysozyme (HEWL) in the presence of dimethyl sulfoxide (DMSO). However, neither binds in aqueous media after 4 d of crystallization and crystal growth, suggesting that DMSO facilitates cisplatin/carboplatin binding to the N atoms of His15 by an unknown mechanism. Crystals of HEWL cocrystallized with cisplatin in both aqueous and DMSO media, of HEWL cocrystallized with carboplatin in DMSO medium and of HEWL cocrystallized with cisplatin and N-acetylglucosamine (NAG) in DMSO medium were stored for between seven and 15 months. X-ray diffraction studies of these crystals were carried out on a Bruker APEX II home-source diffractometer at room temperature. Room-temperature X-ray diffraction data collection removed the need for cryoprotectants to be used, ruling out any effect that the cryoprotectants might have had on binding to the protein. Both cisplatin and carboplatin still bind to both the Nδ and Nℇ atoms of His15 in DMSO media as expected, but more detail for the cyclobutanedicarboxylate (CBDC) moiety of carboplatin was observed at the Nℇ binding site. However, two molecules of cisplatin were now observed to be bound to His15 in aqueous conditions. The platinum peak positions were identified using anomalous difference electron-density maps as a cross-check with Fo-Fc OMIT electron-density maps. The occupancies of each binding site were calculated using SHELXTL. These results show that over time cisplatin binds to both N atoms of His15 of HEWL in aqueous media, whereas this binding is speeded up in the presence of DMSO. The implication of cisplatin binding to proteins after a prolonged period of time is an important consideration for the length of treatment in patients who are given cisplatin.
- School of Chemistry, Faculty of Engineering and Physical Sciences, University Of Manchester, Brunswick Street, Manchester M13 9PL, England.
Organizational Affiliation: