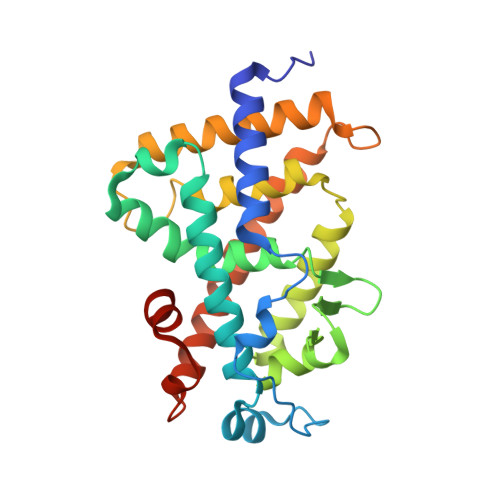
Structural basis for the accommodation of bis- and tris-aromatic derivatives in vitamin d nuclear receptor.
Ciesielski, F., Sato, Y., Chebaro, Y., Moras, D., Dejaegere, A., Rochel, N.(2012) J Med Chem 55: 8440-8449
- PubMed: 22957834 Search on PubMed
- DOI: https://doi.org/10.1021/jm300858s
- Primary Citation Related Structures:
4G1D, 4G1Y, 4G1Z, 4G20, 4G21, 4G2H, 4G2I - PubMed Abstract:
Actual use of the active form of vitamin D (calcitriol or 1α,25-dihydroxyvitamin D(3)) to treat hyperproliferative disorders is hampered by calcemic effects, hence the continuous development of chemically modified analogues with dissociated profiles. Structurally distinct nonsecosteroidal analogues have been developed to mimic calcitriol activity profiles with low calcium serum levels. Here, we report the crystallographic study of vitamin D nuclear receptor (VDR) ligand binding domain in complexes with six nonsecosteroidal analogues harboring two or three phenyl rings. These compounds induce a stimulated transcription in the nanomolar range, similar to calcitriol. Examination of the protein-ligand interactions reveals the mode of binding of these nonsecosteroidal compounds and highlights the role of the various chemical modifications of the ligands to VDR binding and activity, notably (de)solvation effects. The structures with the tris-aromatic ligands exhibit a rearrangement of a novel region of the VDR ligand binding pocket, helix H6.
- Institut de Génétique et de Biologie Moléculaire et Cellulaire (IGBMC), Institut National de Santé et de Recherche Médicale (INSERM) U964, Centre National de Recherche Scientifique (CNRS) UMR 7104, Université de Strasbourg, 67404 Illkirch, France.
Organizational Affiliation: