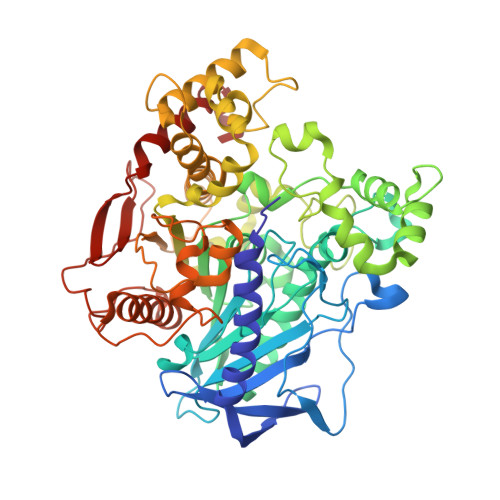
Structure and function of an insect alpha-carboxylesterase ( alpha Esterase7) associated with insecticide resistance.
Jackson, C.J., Liu, J.W., Carr, P.D., Younus, F., Coppin, C., Meirelles, T., Lethier, M., Pandey, G., Ollis, D.L., Russell, R.J., Weik, M., Oakeshott, J.G.(2013) Proc Natl Acad Sci U S A 110: 10177-10182
- PubMed: 23733941 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1304097110
- Primary Citation Related Structures:
4FNG, 4FNM - PubMed Abstract:
Insect carboxylesterases from the αEsterase gene cluster, such as αE7 (also known as E3) from the Australian sheep blowfly Lucilia cuprina (LcαE7), play an important physiological role in lipid metabolism and are implicated in the detoxification of organophosphate (OP) insecticides. Despite the importance of OPs to agriculture and the spread of insect-borne diseases, the molecular basis for the ability of α-carboxylesterases to confer OP resistance to insects is poorly understood. In this work, we used laboratory evolution to increase the thermal stability of LcαE7, allowing its overexpression in Escherichia coli and structure determination. The crystal structure reveals a canonical α/β-hydrolase fold that is very similar to the primary target of OPs (acetylcholinesterase) and a unique N-terminal α-helix that serves as a membrane anchor. Soaking of LcαE7 crystals in OPs led to the capture of a crystallographic snapshot of LcαE7 in its phosphorylated state, which allowed comparison with acetylcholinesterase and rationalization of its ability to protect insects against the effects of OPs. Finally, inspection of the active site of LcαE7 reveals an asymmetric and hydrophobic substrate binding cavity that is well-suited to fatty acid methyl esters, which are hydrolyzed by the enzyme with specificity constants (∼10(6) M(-1) s(-1)) indicative of a natural substrate.
- Research School of Chemistry, Australian National University, Canberra, ACT 0200, Australia. cjackson@rsc.anu.edu.au
Organizational Affiliation: