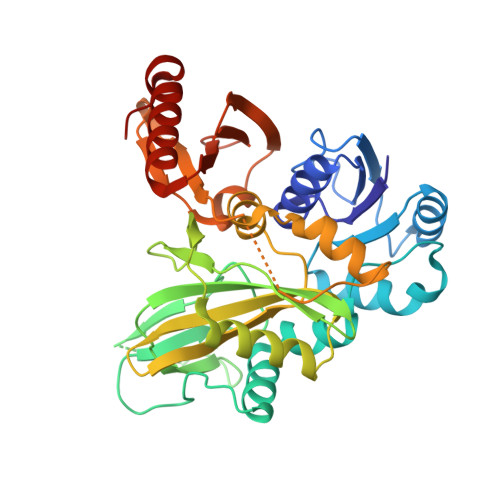
Biosynthesis of the 22nd genetically encoded amino acid pyrrolysine: structure and reaction mechanism of PylC at 1.5A resolution.
Quitterer, F., List, A., Beck, P., Bacher, A., Groll, M.(2012) J Mol Biology 424: 270-282
- PubMed: 22985965 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2012.09.007
- Primary Citation Related Structures:
4FFL, 4FFM, 4FFN, 4FFO, 4FFP, 4FFR - PubMed Abstract:
The second step in the biosynthesis of the 22nd genetically encoded amino acid pyrrolysine (Pyl) is catalyzed by PylC that forms the pseudopeptide L-lysine-N(ε)-3R-methyl-D-ornithine. Here, we present six crystal structures of the monomeric active ligase in complex with substrates, reaction intermediates, and products including ATP, the non-hydrolyzable ATP analogue 5'-adenylyl-β-γ-imidodiphosphate, ADP, D-ornithine (D-Orn), L-lysine (Lys), phosphorylated D-Orn, L-lysine-N(ε)-D-ornithine, inorganic phosphate, carbonate, and Mg(2+). The overall structure of PylC reveals similarities to the superfamily of ATP-grasp enzymes; however, there exist unique structural and functional features for a topological control of successive substrate entry and product release. Furthermore, the presented high-resolution structures provide detailed insights into the reaction mechanism of isopeptide bond formation starting with phosphorylation of D-Orn by transfer of a phosphate moiety from activated ATP. The binding of Lys to the enzyme complex is then followed by an S(N)2 reaction resulting in L-lysine-N(ε)-D-ornithine and inorganic phosphate. Surprisingly, PylC harbors two adenine nucleotides bound at the active site, what has not been observed in any ATP-grasp protein analyzed to date. Whereas one ATP molecule is involved in catalysis, the second adenine nucleotide functions as a selective anchor for the C- and N-terminus of the Lys substrate and is responsible for protein stability as shown by mutagenesis.
- Center for Integrated Protein Science at the Department Chemie, Lehrstuhl für Biochemie, Technische Universität München, Garching D-85747, Germany.
Organizational Affiliation: