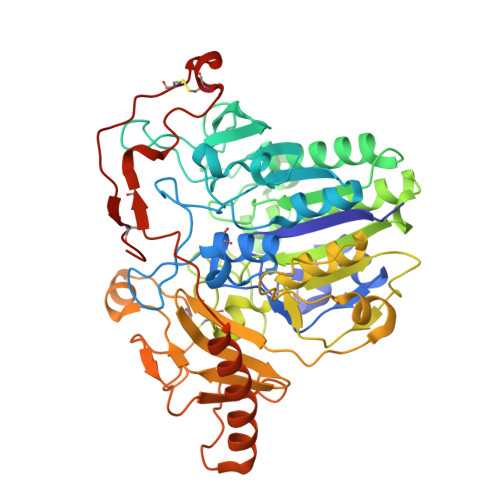
The Structure of Human GALNS Reveals the Molecular Basis for Mucopolysaccharidosis IV A.
Rivera-Colon, Y., Schutsky, E.K., Kita, A.Z., Garman, S.C.(2012) J Mol Biology 423: 736-751
- PubMed: 22940367 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2012.08.020
- Primary Citation Related Structures:
4FDI, 4FDJ - PubMed Abstract:
Lysosomal enzymes catalyze the breakdown of macromolecules in the cell. In humans, loss of activity of a lysosomal enzyme leads to an inherited metabolic defect known as a lysosomal storage disorder. The human lysosomal enzyme galactosamine-6-sulfatase (GALNS, also known as N-acetylgalactosamine-6-sulfatase and GalN6S; E.C. 3.1.6.4) is deficient in patients with the lysosomal storage disease mucopolysaccharidosis IV A (also known as MPS IV A and Morquio A). Here, we report the three-dimensional structure of human GALNS, determined by X-ray crystallography at 2.2Å resolution. The structure reveals a catalytic gem diol nucleophile derived from modification of a cysteine side chain. The active site of GALNS is a large, positively charged trench suitable for binding polyanionic substrates such as keratan sulfate and chondroitin-6-sulfate. Enzymatic assays on the insect-cell-expressed human GALNS indicate activity against synthetic substrates and inhibition by both substrate and product. Mapping 120 MPS IV A missense mutations onto the structure reveals that a majority of mutations affect the hydrophobic core of the structure, indicating that most MPS IV A cases result from misfolding of GALNS. Comparison of the structure of GALNS to paralogous sulfatases shows a wide variety of active-site geometries in the family but strict conservation of the catalytic machinery. Overall, the structure and the known mutations establish the molecular basis for MPS IV A and for the larger MPS family of diseases.
- Department of Biochemistry and Molecular Biology, University of Massachusetts Amherst, Amherst, MA 01003, USA.
Organizational Affiliation: