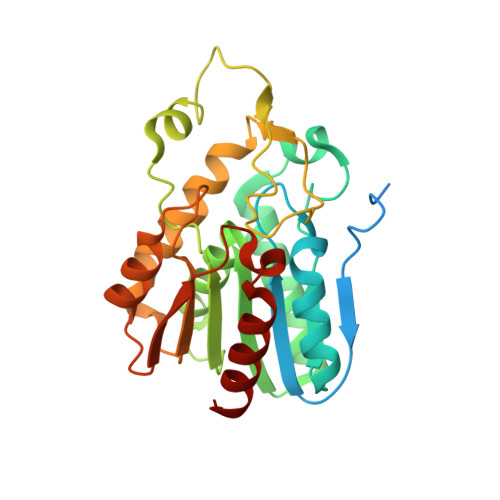
The Metagenome-Derived Enzymes LipS and LipT Increase the Diversity of Known Lipases.
Chow, J., Kovacic, F., Dall Antonia, Y., Krauss, U., Fersini, F., Schmeisser, C., Lauinger, B., Bongen, P., Pietruszka, J., Schmidt, M., Menyes, I., Bornscheuer, U.T., Eckstein, M., Thum, O., Liese, A., Mueller-Dieckmann, J., Jaeger, K.E., Streit, W.R.(2012) PLoS One 7: e47665-e47665
- PubMed: 23112831 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0047665
- Primary Citation Related Structures:
4FBL, 4FBM - PubMed Abstract:
Triacylglycerol lipases (EC 3.1.1.3) catalyze both hydrolysis and synthesis reactions with a broad spectrum of substrates rendering them especially suitable for many biotechnological applications. Most lipases used today originate from mesophilic organisms and are susceptible to thermal denaturation whereas only few possess high thermotolerance. Here, we report on the identification and characterization of two novel thermostable bacterial lipases identified by functional metagenomic screenings. Metagenomic libraries were constructed from enrichment cultures maintained at 65 to 75 °C and screened resulting in the identification of initially 10 clones with lipolytic activities. Subsequently, two ORFs were identified encoding lipases, LipS and LipT. Comparative sequence analyses suggested that both enzymes are members of novel lipase families. LipS is a 30.2 kDa protein and revealed a half-life of 48 h at 70 °C. The lipT gene encoded for a multimeric enzyme with a half-life of 3 h at 70 °C. LipS had an optimum temperature at 70 °C and LipT at 75 °C. Both enzymes catalyzed hydrolysis of long-chain (C(12) and C(14)) fatty acid esters and additionally hydrolyzed a number of industry-relevant substrates. LipS was highly specific for (R)-ibuprofen-phenyl ester with an enantiomeric excess (ee) of 99%. Furthermore, LipS was able to synthesize 1-propyl laurate and 1-tetradecyl myristate at 70 °C with rates similar to those of the lipase CalB from Candida antarctica. LipS represents the first example of a thermostable metagenome-derived lipase with significant synthesis activities. Its X-ray structure was solved with a resolution of 1.99 Å revealing an unusually compact lid structure.
- Department of Microbiology and Biotechnology, Biocenter Klein Flottbek, University of Hamburg, Hamburg, Germany.
Organizational Affiliation: