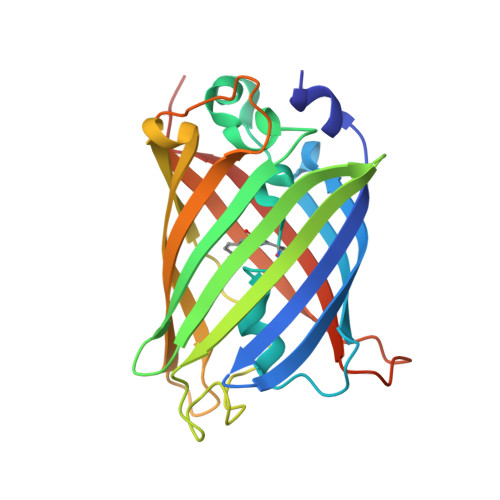
Crystal structure of enhanced green fluorescent protein to 1.35 a resolution reveals alternative conformations for glu222.
Arpino, J.A., Rizkallah, P.J., Jones, D.D.(2012) PLoS One 7: e47132-e47132
- PubMed: 23077555 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0047132
- Primary Citation Related Structures:
4EUL - PubMed Abstract:
Enhanced Green Fluorescent Protein (EGFP) is one of the most widely used engineered variants of the original wild-type Green Fluorescent Protein. Here, we report the high resolution (1.35 Å) structure of EGFP crystallised in its untagged sequence form that reveals the combined impact of the F64L and S65T, that give rise to improved folding and spectral characteristics. The overall structure of EGFP is very similar to wt GFP, forming the classical β-barrel fold with the chromophore containing helix running through the core of the structure. Replacement of Phe64 with Leu in EGFP results in subtle rearrangement of hydrophobic core packing close to the chromophore including the reduction in surface exposure of two hydrophobic residues. Replacement of Ser65 with Thr has a significant impact on the local hydrogen bond network in the vicinity of the chromophore. Detailed analysis of electron density reveals that several residues close to the chromophore occupy at least two distinct conformations. This includes Glu222 that defines the charged state on the chromophore, with the two conformations having slightly different effects on the hydrogen bond network surrounding the chromophore. Hence, the reported high-resolution structure of EGFP has provided a long overdue molecular description of one of the most important fluorescent protein variants currently in general use.
- School of Biosciences, Cardiff University, Cardiff, United Kingdom.
Organizational Affiliation: