The crystal structure of acyl carrier protein phosphodiesterase from Yersinia pestis CO92 in complex with FMN.
Tan, K., Gu, M., Kwon, K., Anderson, W.F., Joachimiak, A.To be published.
Experimental Data Snapshot
Starting Model: experimental
View more details
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| FMN-dependent NADH-azoreductase | 204 | Yersinia pestis CO92 | Mutation(s): 0 Gene Names: azoR, y2010, YPO2323, YP_2110 EC: 1.7 (PDB Primary Data), 1.6.5 (UniProt), 1.7.1.17 (UniProt) | 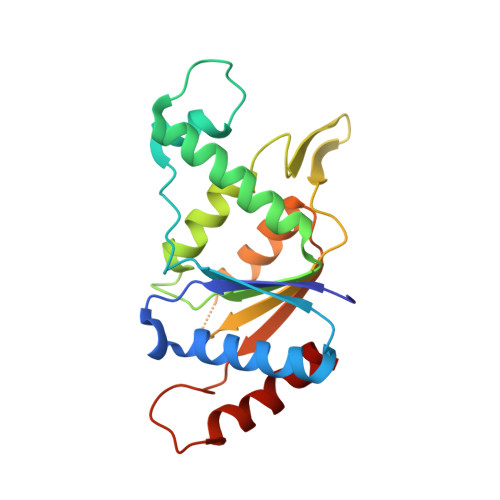 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q8ZE60 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 4 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| 12P Download:Ideal Coordinates CCD File | E [auth A] | DODECAETHYLENE GLYCOL C24 H50 O13 WRZXKWFJEFFURH-UHFFFAOYSA-N |  | ||
| FMN Download:Ideal Coordinates CCD File | B [auth A], C [auth A], D [auth A] | FLAVIN MONONUCLEOTIDE C17 H21 N4 O9 P FVTCRASFADXXNN-SCRDCRAPSA-N |  | ||
| GOL Download:Ideal Coordinates CCD File | F [auth A] | GLYCEROL C3 H8 O3 PEDCQBHIVMGVHV-UHFFFAOYSA-N |  | ||
| IMD Download:Ideal Coordinates CCD File | G [auth A] | IMIDAZOLE C3 H5 N2 RAXXELZNTBOGNW-UHFFFAOYSA-O |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 51.825 | α = 90 |
| b = 120.388 | β = 90 |
| c = 60.074 | γ = 90 |
| Software Name | Purpose |
|---|---|
| SBC-Collect | data collection |
| MOLREP | phasing |
| PHENIX | refinement |
| HKL-3000 | data reduction |
| HKL-3000 | data scaling |