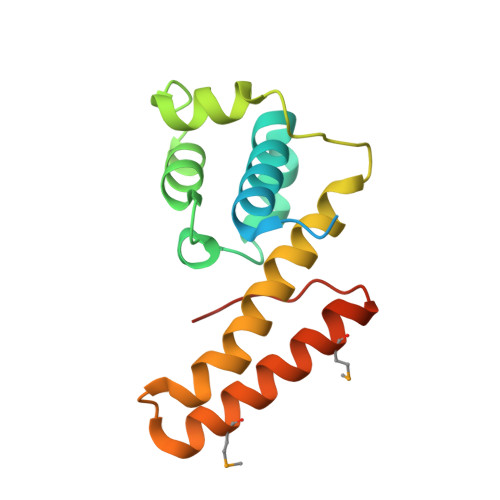
Crystal structure and mode of helicase binding of the C-terminal domain of primase from Helicobacter pylori
Abdul Rehman, S.A., Verma, V., Mazumder, M., Dhar, S.K., Gourinath, S.(2013) J Bacteriol 195: 2826-2838
- PubMed: 23585534
- DOI: https://doi.org/10.1128/JB.00091-13
- Primary Citation Related Structures:
4EHS - PubMed Abstract:
To better understand the poor conservation of the helicase binding domain of primases (DnaGs) among the eubacteria, we determined the crystal structure of the Helicobacter pylori DnaG C-terminal domain (HpDnaG-CTD) at 1.78 Å. The structure has a globular subdomain connected to a helical hairpin. Structural comparison has revealed that globular subdomains, despite the variation in number of helices, have broadly similar arrangements across the species, whereas helical hairpins show different orientations. Further, to study the helicase-primase interaction in H. pylori, a complex was modeled using the HpDnaG-CTD and HpDnaB-NTD (helicase) crystal structures using the Bacillus stearothermophilus BstDnaB-BstDnaG-CTD (helicase-primase) complex structure as a template. By using this model, a nonconserved critical residue Phe534 on helicase binding interface of DnaG-CTD was identified. Mutation guided by molecular dynamics, biophysical, and biochemical studies validated our model. We further concluded that species-specific helicase-primase interactions are influenced by electrostatic surface potentials apart from the critical hydrophobic surface residues.
- School of Life Sciences, Jawaharlal Nehru University, New Delhi, India.
Organizational Affiliation: