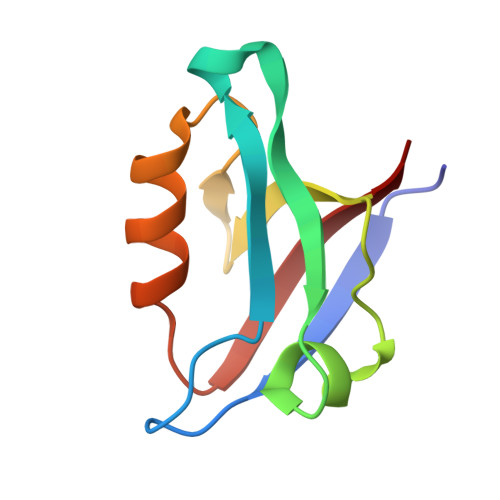
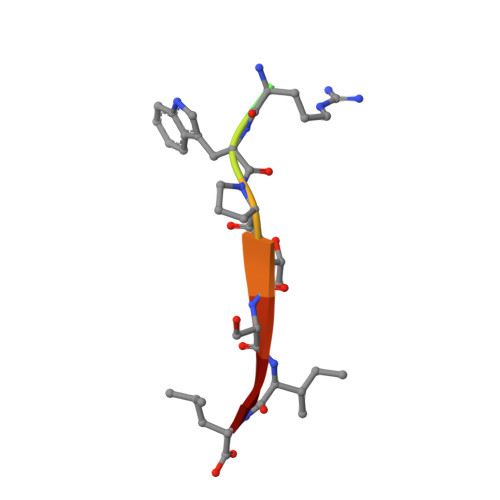
Stereochemical Determinants of C-terminal Specificity in PDZ Peptide-binding Domains: A NOVEL CONTRIBUTION OF THE CARBOXYLATE-BINDING LOOP.
Amacher, J.F., Cushing, P.R., Bahl, C.D., Beck, T., Madden, D.R.(2013) J Biological Chem 288: 5114-5126
- PubMed: 23243314 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M112.401588
- Primary Citation Related Structures:
4E34, 4E35, 4E3B - PubMed Abstract:
PDZ (PSD-95/Dlg/ZO-1) binding domains often serve as cellular traffic engineers, controlling the localization and activity of a wide variety of binding partners. As a result, they play important roles in both physiological and pathological processes. However, PDZ binding specificities overlap, allowing multiple PDZ proteins to mediate distinct effects on shared binding partners. For example, several PDZ domains bind the cystic fibrosis (CF) transmembrane conductance regulator (CFTR), an epithelial ion channel mutated in CF. Among these binding partners, the CFTR-associated ligand (CAL) facilitates post-maturational degradation of the channel and is thus a potential therapeutic target. Using iterative optimization, we previously developed a selective CAL inhibitor peptide (iCAL36). Here, we investigate the stereochemical basis of iCAL36 specificity. The crystal structure of iCAL36 in complex with the CAL PDZ domain reveals stereochemical interactions distributed along the peptide-binding cleft, despite the apparent degeneracy of the CAL binding motif. A critical selectivity determinant that distinguishes CAL from other CFTR-binding PDZ domains is the accommodation of an isoleucine residue at the C-terminal position (P(0)), a characteristic shared with the Tax-interacting protein-1. Comparison of the structures of these two PDZ domains in complex with ligands containing P(0) Leu or Ile residues reveals two distinct modes of accommodation for β-branched C-terminal side chains. Access to each mode is controlled by distinct residues in the carboxylate-binding loop. These studies provide new insights into the primary sequence determinants of binding motifs, which in turn control the scope and evolution of PDZ interactomes.
- Department of Biochemistry, Geisel School of Medicine at Dartmouth, Hanover, New Hampshire 03755, USA.
Organizational Affiliation: