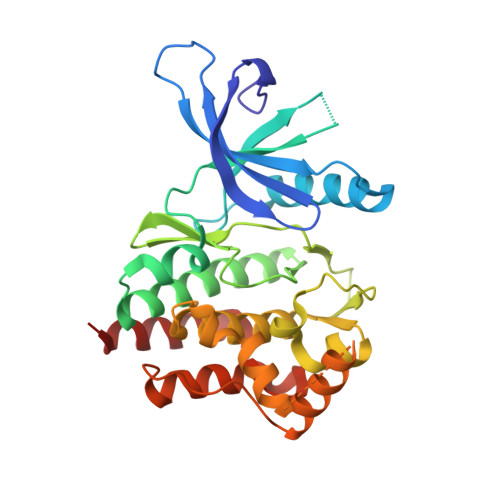
Enabling structure-based drug design of Tyk2 through co-crystallization with a stabilizing aminoindazole inhibitor.
Argiriadi, M.A., Goedken, E.R., Banach, D., Borhani, D.W., Burchat, A., Dixon, R.W., Marcotte, D., Overmeyer, G., Pivorunas, V., Sadhukhan, R., Sousa, S., Moore, N.S., Tomlinson, M., Voss, J., Wang, L., Wishart, N., Woller, K., Talanian, R.V.(2012) BMC Struct Biol 12: 22-22
- PubMed: 22995073 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1186/1472-6807-12-22
- Primary Citation Related Structures:
4E1Z, 4E20 - PubMed Abstract:
Structure-based drug design (SBDD) can accelerate inhibitor lead design and optimization, and efficient methods including protein purification, characterization, crystallization, and high-resolution diffraction are all needed for rapid, iterative structure determination. Janus kinases are important targets that are amenable to structure-based drug design. Here we present the first mouse Tyk2 crystal structures, which are complexed to 3-aminoindazole compounds. A comprehensive construct design effort included N- and C-terminal variations, kinase-inactive mutations, and multiple species orthologs. High-throughput cloning and expression methods were coupled with an abbreviated purification protocol to optimize protein solubility and stability. In total, 50 Tyk2 constructs were generated. Many displayed poor expression, inadequate solubility, or incomplete affinity tag processing. One kinase-inactive murine Tyk2 construct, complexed with an ATP-competitive 3-aminoindazole inhibitor, provided crystals that diffracted to 2.5-2.6 Å resolution. This structure revealed initial "hot-spot" regions for SBDD, and provided a robust platform for ligand soaking experiments. Compared to previously reported human Tyk2 inhibitor crystal structures (Chrencik et al. (2010) J Mol Biol 400:413), our structures revealed a key difference in the glycine-rich loop conformation that is induced by the inhibitor. Ligand binding also conferred resistance to proteolytic degradation by thermolysin. As crystals could not be obtained with the unliganded enzyme, this enhanced stability is likely important for successful crystallization and inhibitor soaking methods. Practical criteria for construct performance and prioritization, the optimization of purification protocols to enhance protein yields and stability, and use of high-throughput construct exploration enable structure determination methods early in the drug discovery process. Additionally, specific ligands stabilize Tyk2 protein and may thereby enable crystallization.
- Department of Molecular & Cellular Pharmacology, Abbott Laboratories, Worcester, MA, USA. maria.argiriadi@abbott.com
Organizational Affiliation: