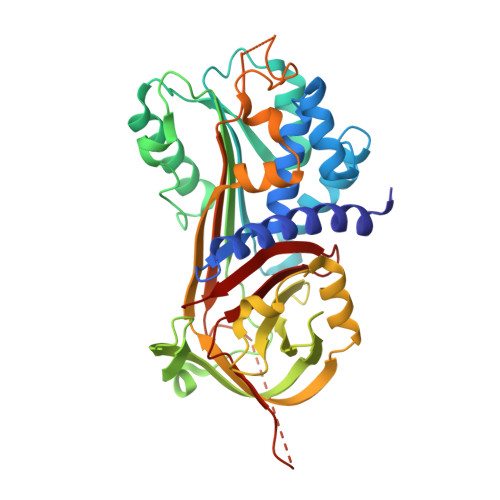
Crystal structures of protease nexin-1 in complex with heparin and thrombin suggest a 2-step recognition mechanism.
Li, W., Huntington, J.A.(2012) Blood 120: 459-467
- PubMed: 22618708 Search on PubMed
- DOI: https://doi.org/10.1182/blood-2012-03-415869
- Primary Citation Related Structures:
4DY0, 4DY7 - PubMed Abstract:
Protease nexin-1 (PN1) is a specific and extremely efficient inhibitor of thrombin. However, unlike other thrombin inhibitors belonging to the serpin family, PN1 is not synthesized in the liver and does not circulate in the blood. Rather, PN1 is expressed by multiple cell types, including macrophages, smooth muscle cells, and platelets, and it is on the surface of these cells, bound to glycosaminoglycans, that PN1 inhibits the signaling functions of thrombin. PN1 sets the threshold for thrombin-induced platelet activation and has been implicated in atherosclerosis. However, in spite of the emerging importance of PN1 in thrombosis and atherosclerosis, little is know about how it associates to cells and how it inhibits thrombin at rates that surpass the diffusion limit. To address these issues, we determined the crystal structures of PN1 in complex with heparin, and in complex with catalytically inert thrombin. The crystal structures suggest a unique 2-step mechanism of thrombin recognition involving rapid electrostatics-driven association to form an initial glycosaminoglycan-bridged complex, followed by a large conformational rearrangement to form the productive Michaelis complex.
- Department of Haematology, Cambridge Institute for Medical Research, University of Cambridge, Addenbrooke's Hospital, Hills Road, Cambridge, United Kingdom. wl225@cam.ac.uk
Organizational Affiliation: