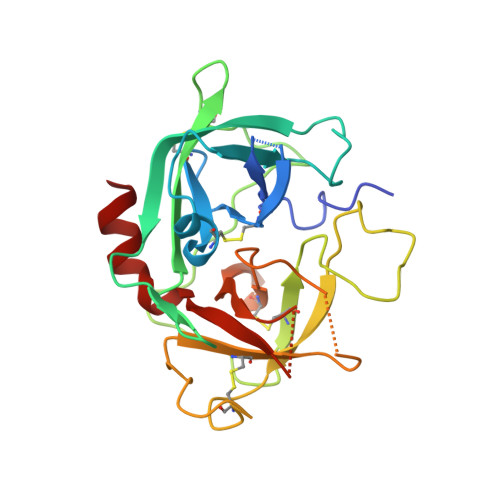
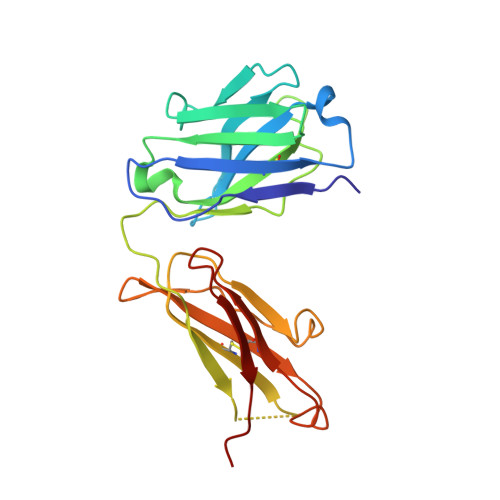
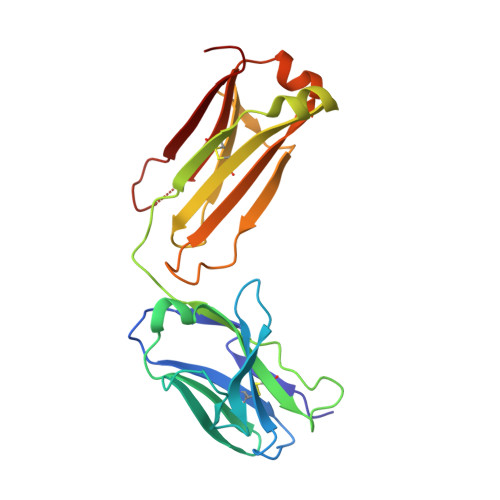
Rezymogenation of active urokinase induced by an inhibitory antibody.
Jiang, L., Botkjaer, K.A., Andersen, L.M., Yuan, C., Andreasen, P.A., Huang, M.(2013) Biochem J 449: 161-166
- PubMed: 23016918 Search on PubMed
- DOI: https://doi.org/10.1042/BJ20121132
- Primary Citation Related Structures:
4DVA, 4DVB, 4DW2 - PubMed Abstract:
An important regulatory mechanism of serine proteases is the proteolytic conversion of the inactive pro-enzyme, or zymogen, into the active enzyme. This activation process is generally considered an irreversible process. In the present study, we demonstrate that an active enzyme can be converted back into its zymogen form. We determined the crystal structure of uPA (urokinase-type plasminogen activator) in complex with an inhibitory antibody, revealing that the antibody 'rezymogenizes' already activated uPA. The present study demonstrates a new regulatory mechanism of protease activity, which is also an extreme case of protein allostery. Mechanistically, the antibody binds a single surface-exposed loop, named the autolysis loop, thereby preventing the stabilization of uPA in its active conformation. We argue that this autolysis loop is a key structural element for rezymogenation of other proteases, and will be a new target site for pharmacological intervention with serine protease activity.
- State Key Laboratory of Structural Chemistry, Fujian Institute of Research on the Structure of Matter, Chinese Academy of Sciences, 155 Yang Qiao West Road, Fuzhou, Fujian, 350002, China.
Organizational Affiliation: