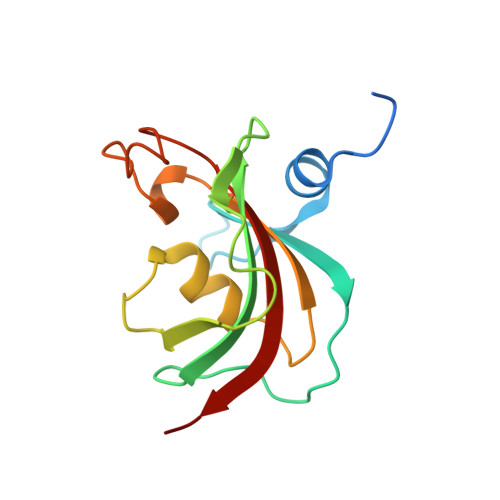
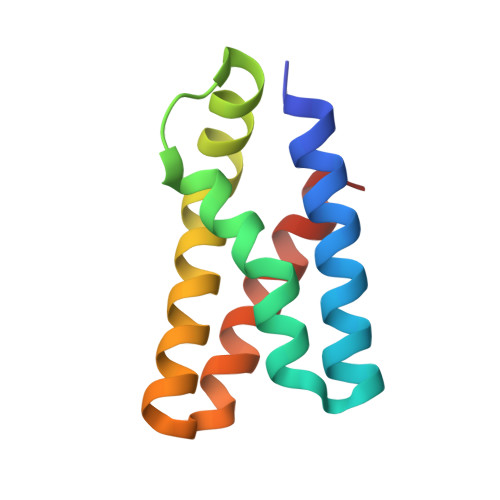
Large FK506-Binding Proteins Shape the Pharmacology of Rapamycin.
Marz, A.M., Fabian, A.K., Kozany, C., Bracher, A., Hausch, F.(2013) Mol Cell Biol 33: 1357-1367
- PubMed: 23358420 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/MCB.00678-12
- Primary Citation Related Structures:
4DRH, 4DRI, 4DRJ - PubMed Abstract:
The immunosuppressant and anticancer drug rapamycin works by inducing inhibitory protein complexes with the kinase mTOR, an important regulator of growth and proliferation. The obligatory accessory partner of rapamycin is believed to be FK506-binding protein 12 (FKBP12). Here we show that rapamycin complexes of larger FKBP family members can tightly bind to mTOR and potently inhibit its kinase activity. Cocrystal structures with FKBP51 and FKBP52 reveal the modified molecular binding mode of these alternative ternary complexes in detail. In cellular model systems, FKBP12 can be functionally replaced by larger FKBPs. When the rapamycin dosage is limiting, mTOR inhibition of S6K phosphorylation can be enhanced by FKBP51 overexpression in mammalian cells, whereas FKBP12 is dispensable. FKBP51 could also enable the rapamycin-induced hyperphosphorylation of Akt, which depended on higher FKBP levels than rapamycin-induced inhibition of S6K phosphorylation. These insights provide a mechanistic rationale for preferential mTOR inhibition in specific cell or tissue types by engaging specific FKBP homologs.
- Max Planck Institute of Psychiatry, Department of Chemical Genomics, Munich, Germany.
Organizational Affiliation: