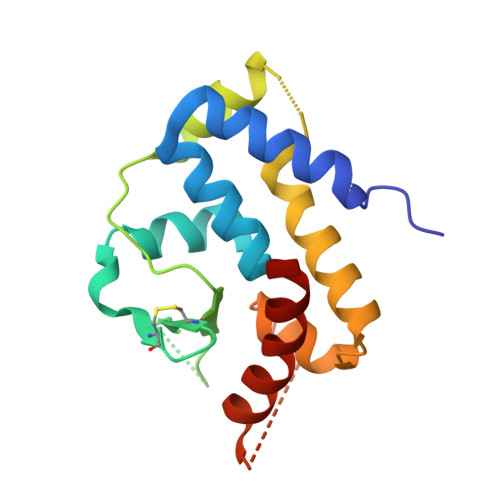
Structural and functional characterization of the C-terminal catalytic domain of SSV1 integrase.
Zhan, Z., Ouyang, S., Liang, W., Zhang, Z., Liu, Z.J., Huang, L.(2012) Acta Crystallogr D Biol Crystallogr 68: 659-670
- PubMed: 22683788 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444912007202
- Primary Citation Related Structures:
3VCF, 4DKS - PubMed Abstract:
The spindle-shaped virus SSV1 of the hyperthermophilic archaeon Sulfolobus shibatae encodes an integrase (SSV1 Int). Here, the crystal structure of the C-terminal catalytic domain of SSV1 Int is reported. This is the first structural study of an archaeal tyrosine recombinase. Structural comparison shows that the C-terminal domain of SSV1 Int possesses a core fold similar to those of tyrosine recombinases of both bacterial and eukaryal origin, apart from the lack of a conserved helix corresponding to αI of Cre, indicating conservation of these enzymes among all three domains of life. Five of the six catalytic residues cluster around a basic cleft on the surface of the structure and the nucleophile Tyr314 is located on a flexible loop that stretches away from the central cleft, supporting the possibility that SSV1 Int cleaves the target DNA in a trans mode. Biochemical analysis suggests that the N-terminal domain is responsible for the dimerization of SSV1 Int. The C-terminal domain is capable of DNA cleavage and ligation, but at efficiencies significantly lower than those of the full-length protein. In addition, neither the N-terminal domain alone nor the C-terminal domain alone shows a strong sequence preference in DNA binding. Therefore, recognition of the core-type sequence and efficient catalysis by SSV1 Int presumably requires covalent linkage and interdomain communication between the two domains.
- State Key Laboratory of Microbial Resources, Institute of Microbiology, Chinese Academy of Sciences, No. 1 West Beichen Road, Chaoyang District, Beijing 100101, People's Republic of China.
Organizational Affiliation: