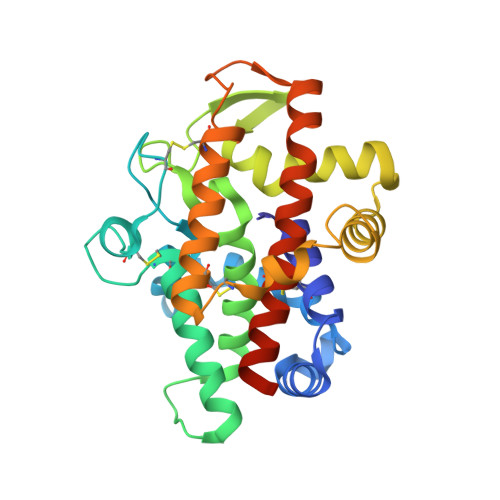
Plant multifunctional nuclease TBN1 with unexpected phospholipase activity: structural study and reaction-mechanism analysis.
Koval, T., Lipovova, P., Podzimek, T., Matousek, J., Duskova, J., Skalova, T., Stepankova, A., Hasek, J., Dohnalek, J.(2013) Acta Crystallogr D Biol Crystallogr 69: 213-226
- PubMed: 23385457 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444912043697
- Primary Citation Related Structures:
3SNG, 4DJ4 - PubMed Abstract:
Type I plant nucleases play an important role in apoptotic processes and cell senescence. Recently, they have also been indicated to be potent anticancer agents in in vivo studies. The first structure of tomato nuclease I (TBN1) has been determined, its oligomerization and activity profiles have been analyzed and its unexpected activity towards phospholipids has been discovered, and conclusions are drawn regarding its catalytic mechanism. The structure-solution process required X-ray diffraction data from two crystal forms. The first form was used for phase determination; the second form was used for model building and refinement. TBN1 is mainly α-helical and is stabilized by four disulfide bridges. Three observed oligosaccharides are crucial for its stability and solubility. The active site is localized at the bottom of the positively charged groove and contains a zinc cluster that is essential for enzymatic activity. An equilibrium between monomers, dimers and higher oligomers of TBN1 was observed in solution. Principles of the reaction mechanism of the phosphodiesterase activity are suggested, with central roles for the zinc cluster, the nucleobase-binding pocket (Phe-site) and Asp70, Arg73 and Asn167. Based on the distribution of surface residues, possible binding sites for dsDNA and other nucleic acids with secondary structure were identified. The phospholipase activity of TBN1, which is reported for the first time for a nuclease, significantly broadens the substrate promiscuity of the enzyme, and the resulting release of diacylglycerol, which is an important second messenger, can be related to the role of TBN1 in apoptosis.
- Institute of Macromolecular Chemistry, AS CR, v.v.i., Heyrovskeho nam. 2, 162 06 Praha 6, Czech Republic. koval.tomas@gmail.com
Organizational Affiliation: