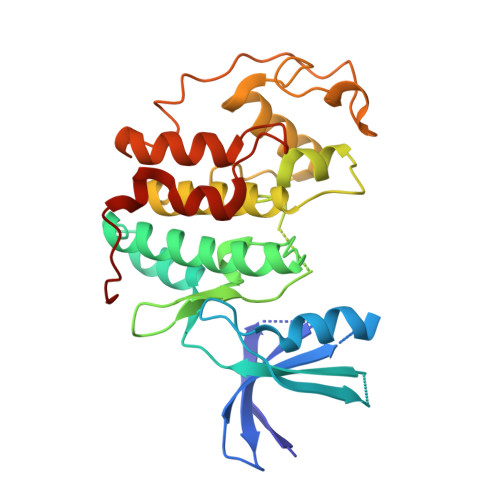
Luciferin and Derivatives as a Dyrk Selective Scaffold for the Design of Protein Kinase Inhibitors.
Rothweiler, U., Eriksson, J., Stensen, W., Leeson, F., Engh, R.A., Svendsen, J.S.(2015) Eur J Med Chem 94: 140
- PubMed: 25768698 Search on PubMed
- DOI: https://doi.org/10.1016/j.ejmech.2015.02.035
- Primary Citation Related Structures:
4D1X, 4D1Z - PubMed Abstract:
D-Luciferin is widely used as a substrate in luciferase catalysed bioluminescence assays for in vitro studies. However, little is known about cross reactivity and potential interference of D-luciferin with other enzymes. We serendipitously found that firefly luciferin inhibited the CDK2/Cyclin A protein kinase. Inhibition profiling of D-luciferin over a 103-protein kinase panel showed significant inhibition of a small set of protein kinases, in particular the DYRK-family, but also other members of the CMGC-group, including ERK8 and CK2. Inhibition profiling on a 16-member focused library derived from D-luciferin confirms that D-luciferin represents a DYRK-selective chemotype of fragment-like molecular weight. Thus, observation of its inhibitory activity and the initial SAR information reported here promise to be useful for further design of protein kinase inhibitors with related scaffolds.
- The Norwegian Structural Biology Centre, Department of Chemistry, UiT The Arctic University of Norway, N-9037 Tromsø, Norway.
Organizational Affiliation: