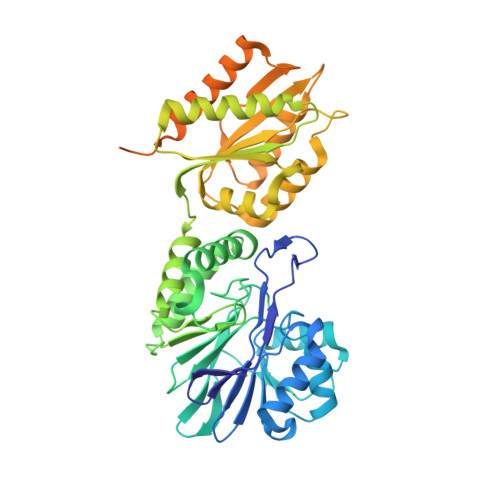
Structure of Escherichia Coli Flavodiiron Nitric Oxide Reductase.
Romao, C.V., Vicente, J.B., Borges, P.T., Victor, B.L., Lamosa, P., Silva, E., Bandeiras, T.M., Soares, C.M., Carrondo, M.A., Turner, D., Teixeira, M., Frazao, C.(2016) J Mol Biology 428: 4686
- PubMed: 27725182 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2016.10.008
- Primary Citation Related Structures:
4D02, 5LLD, 5LMC - PubMed Abstract:
Flavodiiron proteins (FDPs) are present in organisms from all domains of life and have been described so far to be involved in the detoxification of oxygen or nitric oxide (NO), acting as O 2 and/or NO reductases. The Escherichia coli FDP, named flavorubredoxin (FlRd), is the most extensively studied FDP. Biochemical and in vivo studies revealed that FlRd is involved in NO detoxification as part of the bacterial defense mechanisms against reactive nitrogen species. E. coli FlRd has a clear preference for NO as a substrate in vitro, exhibiting a very low reactivity toward O 2 . To contribute to the understanding of the structural features defining this substrate selectivity, we determined the crystallographic structure of E. coli FlRd, both in the isolated and reduced states. The overall tetrameric structure revealed a highly conserved flavodiiron core domain, with a metallo-β-lactamase-like domain containing a diiron center, and a flavodoxin domain with a flavin mononucleotide cofactor. The metal center in the oxidized state has a μ-hydroxo bridge coordinating the two irons, while in the reduced state, this moiety is not detected. Since only the flavodiiron domain was observed in these crystal structures, the structure of the rubredoxin domain was determined by NMR. Tunnels for the substrates were identified, and through molecular dynamics simulations, no differences for O 2 or NO permeation were found. The present data represent the first structure for a NO-selective FDP.
- Instituto de Tecnologia Química e Biológica António Xavier, ITQB NOVA, Av. da República, 2780-157 Oeiras, Portugal.
Organizational Affiliation: