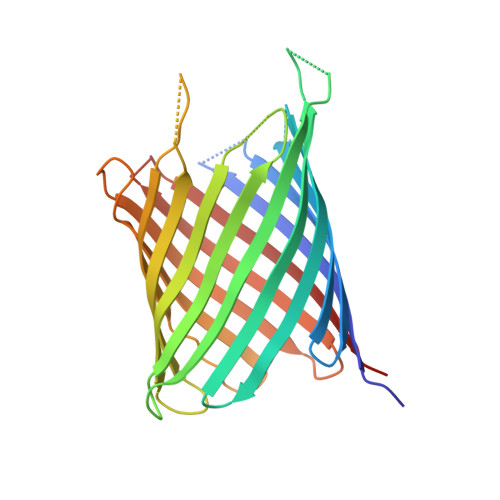
Structure-Based Engineering of a Minimal Porin Reveals Loop- Independent Channel Closure.
Grosse, W., Psakis, G., Mertins, B., Reiss, P., Windisch, D., Brademann, F., Burck, J., Ulrich, A., Koert, U., Essen, L.(2014) Biochemistry 53: 4826
- PubMed: 24988371 Search on PubMed
- DOI: https://doi.org/10.1021/bi500660q
- Primary Citation Related Structures:
4CTD - PubMed Abstract:
Porins, like outer membrane protein G (OmpG) of Escherichia coli, are ideal templates among ion channels for protein and chemical engineering because of their robustness and simple architecture. OmpG shows fast transitions between open and closed states, which were attributed to loop 6 (L6). As flickering limits single-channel-based applications, we pruned L6 by either 8 or 12 amino acids. While the open probabilities of both L6 variants resemble that of native OmpG, their gating frequencies were reduced by 63 and 81%, respectively. Using the 3.2 Å structure of the shorter L6 variant in the open state, we engineered a minimal porin (220 amino acids), where all remaining extramembranous loops were truncated. Unexpectedly, this minimized porin still exhibited gating, but it was 5-fold less frequent than in OmpG. The residual gating of the minimal pore is hence independent of L6 rearrangements and involves narrowing of the ion conductance pathway most probably driven by global stretching-flexing deformations of the membrane-embedded β-barrel.
- Department of Chemistry, Philipps-University Marburg , Hans-Meerwein-Straße, 35032 Marburg, Germany.
Organizational Affiliation: