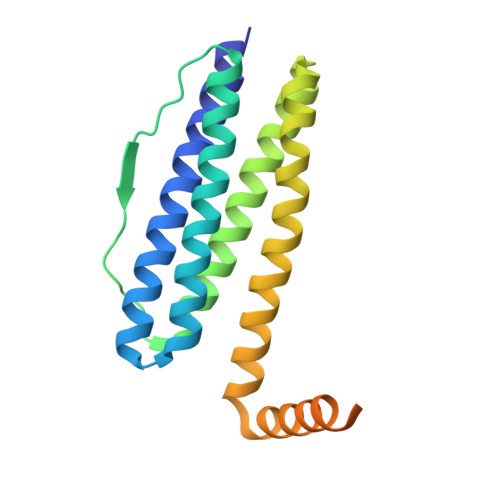
The Crystal Structure of Ferritin from Chlorobium Tepidum Reveals a New Conformation of the 4-Fold Channel for This Protein Family.
Arenas, M., Townsend, P.D., Brito, C., Marquez, V., Marabolli, V., Gonzalez-Nilo, F., Matias, C., Watt, R.K., Lopez-Castro, J.D., Dominguez-Vera, J., Pohl, E., Yevenes, A.(2014) Biochimie 106: 39
- PubMed: 25079050 Search on PubMed
- DOI: https://doi.org/10.1016/j.biochi.2014.07.019
- Primary Citation Related Structures:
4CMY - PubMed Abstract:
Ferritins are ubiquitous iron-storage proteins found in all kingdoms of life. They share a common architecture made of 24 subunits of five α-helices. The recombinant Chlorobium tepidum ferritin (rCtFtn) is a structurally interesting protein since sequence alignments with other ferritins show that this protein has a significantly extended C-terminus, which possesses 12 histidine residues as well as several aspartate and glutamic acid residues that are potential metal ion binding residues. We show that the macromolecular assembly of rCtFtn exhibits a cage-like hollow shell consisting of 24 monomers that are related by 4-3-2 symmetry; similar to the assembly of other ferritins. In all ferritins of known structure the short fifth α-helix adopts an acute angle with respect to the four-helix bundle. However, the crystal structure of the rCtFtn presented here shows that this helix adopts a new conformation defining a new assembly of the 4-fold channel of rCtFtn. This conformation allows the arrangement of the C-terminal region into the inner cavity of the protein shell. Furthermore, two Fe(III) ions were found in each ferroxidase center of rCtFtn, with an average FeA-FeB distance of 3 Å; corresponding to a diferric μ-oxo/hydroxo species. This is the first ferritin crystal structure with an isolated di-iron center in an iron-storage ferritin. The crystal structure of rCtFtn and the biochemical results presented here, suggests that rCtFtn presents similar biochemical properties reported for other members of this protein family albeit with distinct structural plasticity.
- Centro de Bioinformática y Simulación Molecular, Facultad de Ingeniería, Universidad de Talca, Talca, Chile.
Organizational Affiliation: