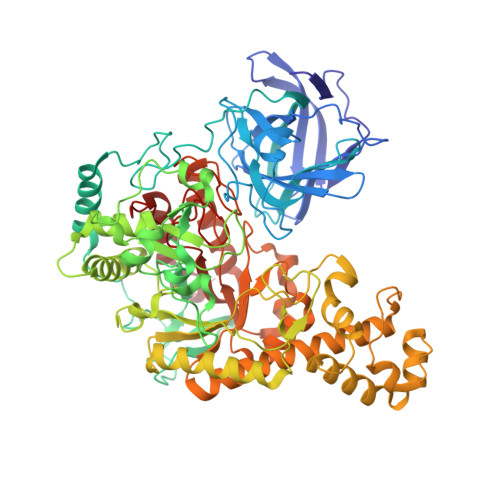
Substrate Recognition and Hydrolysis by a Family 50 Exo-Beta-Agarase Aga50D from the Marine Bacterium Saccharophagus Degradans
Pluvinage, B., Hehemann, J.H., Boraston, A.B.(2013) J Biological Chem 288: 28078
- PubMed: 23921382 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M113.491068
- Primary Citation Related Structures:
4BQ2, 4BQ3, 4BQ4, 4BQ5 - PubMed Abstract:
The bacteria that metabolize agarose use multiple enzymes of complementary specificities to hydrolyze the glycosidic linkages in agarose, a linear polymer comprising the repeating disaccharide subunit of neoagarobiose (3,6-anhydro-l-galactose-α-(1,3)-d-galactose) that are β-(1,4)-linked. Here we present the crystal structure of a glycoside hydrolase family 50 exo-β-agarase, Aga50D, from the marine microbe Saccharophagus degradans. This enzyme catalyzes a critical step in the metabolism of agarose by S. degradans through cleaving agarose oligomers into neoagarobiose products that can be further processed into monomers. The crystal structure of Aga50D to 1.9 Å resolution reveals a (β/α)8-barrel fold that is elaborated with a β-sandwich domain and extensive loops. The structures of catalytically inactivated Aga50D in complex with non-hydrolyzed neoagarotetraose (2.05 Å resolution) and neoagarooctaose (2.30 Å resolution) provide views of Michaelis complexes for a β-agarase. In these structures, the d-galactose residue in the -1 subsite is distorted into a (1)S3 skew boat conformation. The relative positioning of the putative catalytic residues are most consistent with a retaining catalytic mechanism. Additionally, the neoagarooctaose complex showed that this extended substrate made substantial interactions with the β-sandwich domain, which resembles a carbohydrate-binding module, thus creating additional plus (+) subsites and funneling the polymeric substrate through the tunnel-shaped active site. A synthesis of these results in combination with an additional neoagarobiose product complex suggests a potential exo-processive mode of action of Aga50D on the agarose double helix.
- From the Department of Biochemistry and Microbiology, University of Victoria, Victoria, British Columbia V8W 3P6, Canada and.
Organizational Affiliation: