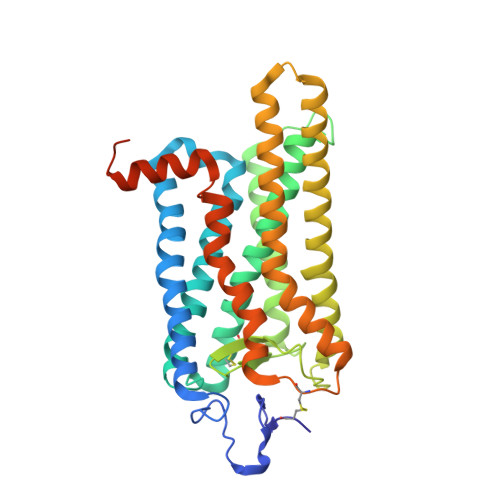
Insights Into Congenital Stationary Night Blindness Based on the Structure of G90D Rhodopsin.
Singhal, A., Ostermaier, M.K., Vishnivetskiy, S.A., Panneels, V., Homan, K.T., Tesmer, J.J., Veprintsev, D., Deupi, X., Gurevich, V.V., Schertler, G.F., Standfuss, J.(2013) EMBO Rep 14: 520
- PubMed: 23579341 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/embor.2013.44
- Primary Citation Related Structures:
4BEY, 4BEZ - PubMed Abstract:
We present active-state structures of the G protein-coupled receptor (GPCRs) rhodopsin carrying the disease-causing mutation G90D. Mutations of G90 cause either retinitis pigmentosa (RP) or congenital stationary night blindness (CSNB), a milder, non-progressive form of RP. Our analysis shows that the CSNB-causing G90D mutation introduces a salt bridge with K296. The mutant thus interferes with the E113Q-K296 activation switch and the covalent binding of the inverse agonist 11-cis-retinal, two interactions that are crucial for the deactivation of rhodopsin. Other mutations, including G90V causing RP, cannot promote similar interactions. We discuss our findings in context of a model in which CSNB is caused by constitutive activation of the visual signalling cascade.
- Laboratory of Biomolecular Research, Paul Scherrer Institut, Villigen 5232, Switzerland.
Organizational Affiliation: