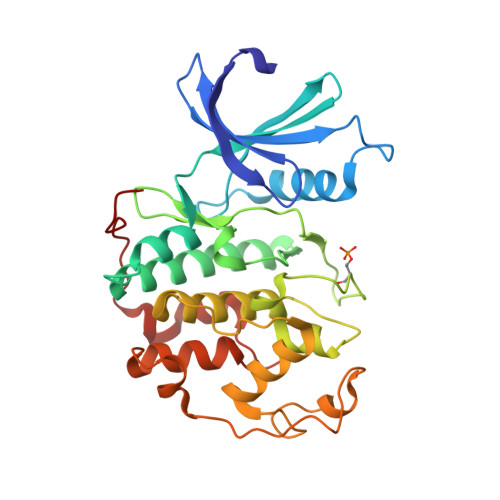
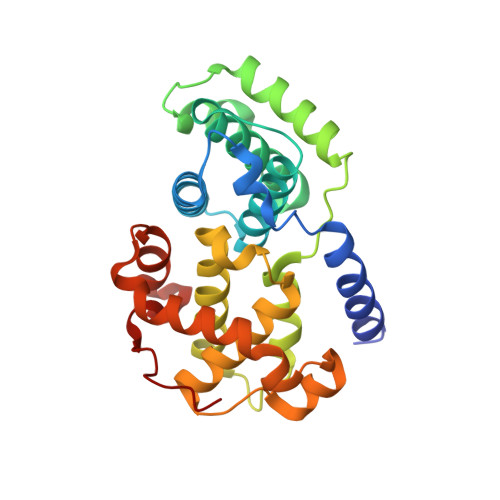
Substituted 4-(Thiazol-5-Yl)-2-(Phenylamino)Pyrimidines are Highly Active Cdk9 Inhibitors: Synthesis, X-Ray Crystal Structure, Sar and Anti-Cancer Activities.
Shao, H., Shi, S., Huang, S., Hole, A., Abbas, A.Y., Baumli, S., Liu, X., Lam, F., Foley, D.W., Fischer, P.M., Noble, M., Endicott, J.A., Pepper, C., Wang, S.(2013) J Med Chem 56: 640
- PubMed: 23301767 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/jm301475f
- Primary Citation Related Structures:
4BCG, 4BCP - PubMed Abstract:
Cancer cells often have a high demand for antiapoptotic proteins in order to resist programmed cell death. CDK9 inhibition selectively targets survival proteins and reinstates apoptosis in cancer cells. We designed a series of 4-thiazol-2-anilinopyrimidine derivatives with functional groups attached to the C5-position of the pyrimidine or to the C4-thiazol moiety and investigated their effects on CDK9 potency and selectivity. One of the most selective compounds, 12u inhibits CDK9 with IC(50) = 7 nM and shows over 80-fold selectivity for CDK9 versus CDK2. X-ray crystal structures of 12u bound to CDK9 and CDK2 provide insights into the binding modes. This work, together with crystal structures of selected inhibitors in complex with both enzymes described in a companion paper, (34) provides a rationale for the observed SAR. 12u demonstrates potent anticancer activity against primary chronic lymphocytic leukemia cells with a therapeutic window 31- and 107-fold over those of normal B- and T-cells.
- School of Pharmacy and Centre for Biomolecular Sciences, University of Nottingham, University Park, Nottingham NG7 2RD, UK.
Organizational Affiliation: