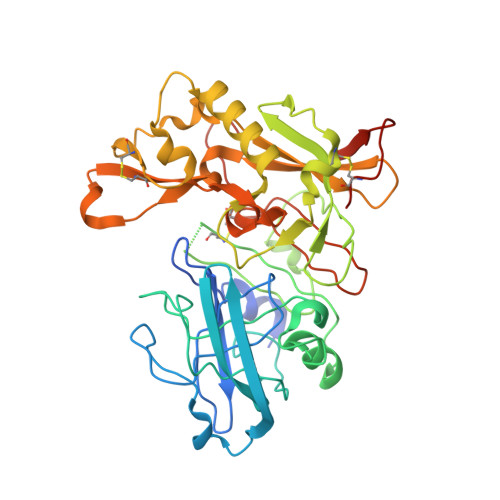
Design and synthesis of beta-site amyloid precursor protein cleaving enzyme (BACE1) inhibitors with in vivo brain reduction of beta-amyloid peptides.
Swahn, B.M., Kolmodin, K., Karlstrom, S., von Berg, S., Soderman, P., Holenz, J., Berg, S., Lindstrom, J., Sundstrom, M., Turek, D., Kihlstrom, J., Slivo, C., Andersson, L., Pyring, D., Rotticci, D., Ohberg, L., Kers, A., Bogar, K., von Kieseritzky, F., Bergh, M., Olsson, L.L., Janson, J., Eketjall, S., Georgievska, B., Jeppsson, F., Falting, J.(2012) J Med Chem 55: 9346-9361
- PubMed: 22924815 Search on PubMed
- DOI: https://doi.org/10.1021/jm3009025
- Primary Citation Related Structures:
4AZY, 4B00 - PubMed Abstract:
The evaluation of a series of aminoisoindoles as β-site amyloid precursor protein cleaving enzyme 1 (BACE1) inhibitors and the discovery of a clinical candidate drug for Alzheimer's disease, (S)-32 (AZD3839), are described. The improvement in permeability properties by the introduction of fluorine adjacent to the amidine moiety, resulting in in vivo brain reduction of Aβ40, is discussed. Due to the basic nature of these compounds, they displayed affinity for the human ether-a-go-go related gene (hERG) ion channel. Different ways to reduce hERG inhibition and increase hERG margins for this series are described, culminating in (S)-16 and (R)-41 showing large in vitro margins with BACE1 cell IC(50) values of 8.6 and 0.16 nM, respectively, and hERG IC(50) values of 16 and 2.8 μM, respectively. Several compounds were advanced into pharmacodynamic studies and demonstrated significant reduction of β-amyloid peptides in mouse brain following oral dosing.
- Department of Medicinal Chemistry, AstraZeneca R&D Södertälje, SE-151 85, Södertälje, Sweden. brittmarieswahn@gmail.com
Organizational Affiliation: