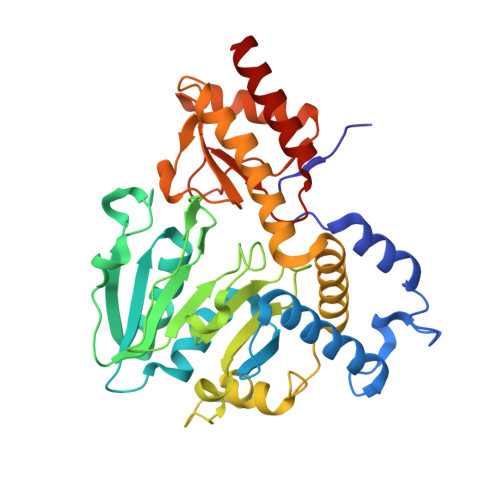
Structural Basis of L-Phosphoserine Binding to Bacillus Alcalophilus Phosphoserine Aminotransferase
Battula, P., Dubnovitsky, A.P., Papageorgiou, A.C.(2013) Acta Crystallogr D Biol Crystallogr 69: 804
- PubMed: 23633589 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444913002096
- Primary Citation Related Structures:
4AZJ, 4AZK - PubMed Abstract:
Phosphoserine aminotransferase is a vitamin B6-dependent enzyme that catalyzes the reversible conversion of 3-phosphohydroxypyruvate to L-phosphoserine using glutamate as an amine donor. In an effort to gain insight into the substrate-recognition mechanism of the enzyme, crystal structures of Bacillus alcalophilus phosphoserine aminotransferase in the presence or absence of L-phosphoserine were determined to resolutions of 1.5 and 1.6 Å, respectively. Local conformational changes induced upon substrate binding were identified. However, in contrast to other aminotransferases, no domain or subunit movements were observed. Two Arg residues (Arg42 and Arg328) and two His residues (His41 and His327) were found to form a tight binding site for the phosphate group of L-phosphoserine. Comparison with Escherichia coli phosphoserine aminotransferase in complex with the substrate analogue α-methylglutamate revealed more extensive structural changes in the case of L-phosphoserine binding. Based on the structural analysis, the flexibility of Arg328 is proposed to be critical for substrate recognition.
- Turku Centre for Biotechnology, University of Turku and Åbo Akademi University, BioCity, FIN-20521 Turku, Finland.
Organizational Affiliation: