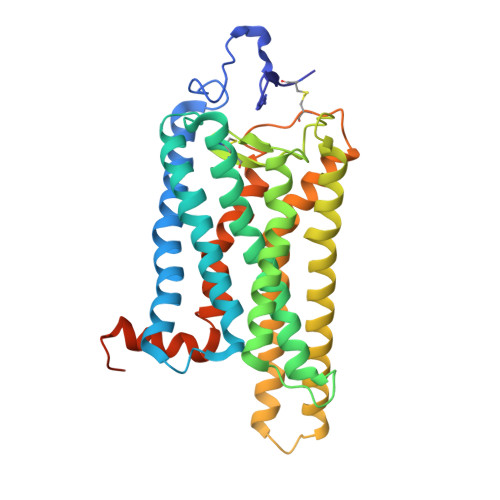
Stabilized G Protein Binding Site in the Structure of Constitutively Active Metarhodopsin-II.
Deupi, X., Edwards, P., Singhal, A., Nickle, B., Oprian, D., Schertler, G., Standfuss, J.(2012) Proc Natl Acad Sci U S A 109: 119
- PubMed: 22198838 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1114089108
- Primary Citation Related Structures:
4A4M - PubMed Abstract:
G protein-coupled receptors (GPCR) are seven transmembrane helix proteins that couple binding of extracellular ligands to conformational changes and activation of intracellular G proteins, GPCR kinases, and arrestins. Constitutively active mutants are ubiquitously found among GPCRs and increase the inherent basal activity of the receptor, which often correlates with a pathological outcome. Here, we have used the M257Y(6.40) constitutively active mutant of the photoreceptor rhodopsin in combination with the specific binding of a C-terminal fragment from the G protein alpha subunit (GαCT) to trap a light activated state for crystallization. The structure of the M257Y/GαCT complex contains the agonist all-trans-retinal covalently bound to the native binding pocket and resembles the G protein binding metarhodopsin-II conformation obtained by the natural activation mechanism; i.e., illumination of the prebound chromophore 11-cis-retinal. The structure further suggests a molecular basis for the constitutive activity of 6.40 substitutions and the strong effect of the introduced tyrosine based on specific interactions with Y223(5.58) in helix 5, Y306(7.53) of the NPxxY motif and R135(3.50) of the E(D)RY motif, highly conserved residues of the G protein binding site.
- Paul Scherrer Institut, Villigen PSI, 5232, Switzerland.
Organizational Affiliation: