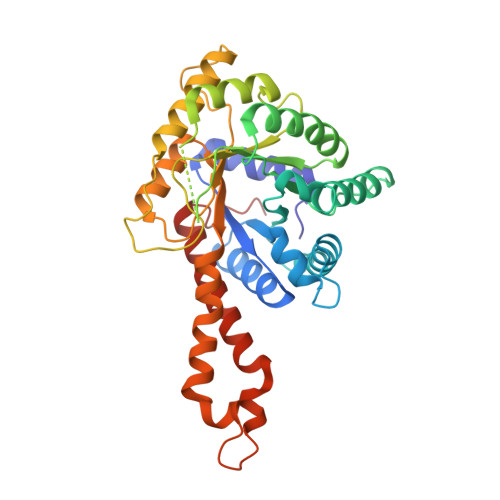
Glycolytic and non-glycolytic functions of Mycobacterium tuberculosis fructose-1,6-bisphosphate aldolase, an essential enzyme produced by replicating and non-replicating bacilli.
de la Paz Santangelo, M., Gest, P.M., Guerin, M.E., Coincon, M., Pham, H., Ryan, G., Puckett, S.E., Spencer, J.S., Gonzalez-Juarrero, M., Daher, R., Lenaerts, A.J., Schnappinger, D., Therisod, M., Ehrt, S., Sygusch, J., Jackson, M.(2011) J Biological Chem 286: 40219-40231
- PubMed: 21949126 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M111.259440
- Primary Citation Related Structures:
4A21, 4A22 - PubMed Abstract:
The search for antituberculosis drugs active against persistent bacilli has led to our interest in metallodependent class II fructose-1,6-bisphosphate aldolase (FBA-tb), a key enzyme of gluconeogenesis absent from mammalian cells. Knock-out experiments at the fba-tb locus indicated that this gene is required for the growth of Mycobacterium tuberculosis on gluconeogenetic substrates and in glucose-containing medium. Surface labeling and enzymatic activity measurements revealed that this enzyme was exported to the cell surface of M. tuberculosis and produced under various axenic growth conditions including oxygen depletion and hence by non-replicating bacilli. Importantly, FBA-tb was also produced in vivo in the lungs of infected guinea pigs and mice. FBA-tb bound human plasmin(ogen) and protected FBA-tb-bound plasmin from regulation by α(2)-antiplasmin, suggestive of an involvement of this enzyme in host/pathogen interactions. The crystal structures of FBA-tb in the native form and in complex with a hydroxamate substrate analog were determined to 2.35- and 1.9-Å resolution, respectively. Whereas inhibitor attachment had no effect on the plasminogen binding activity of FBA-tb, it competed with the natural substrate of the enzyme, fructose 1,6-bisphosphate, and substantiated a previously unknown reaction mechanism associated with metallodependent aldolases involving recruitment of the catalytic zinc ion by the substrate upon active site binding. Altogether, our results highlight the potential of FBA-tb as a novel therapeutic target against both replicating and non-replicating bacilli.
- Mycobacteria Research Laboratories, Department of Microbiology, Immunology, and Pathology, Colorado State University, Fort Collins, Colorado 80523-1682, USA.
Organizational Affiliation: