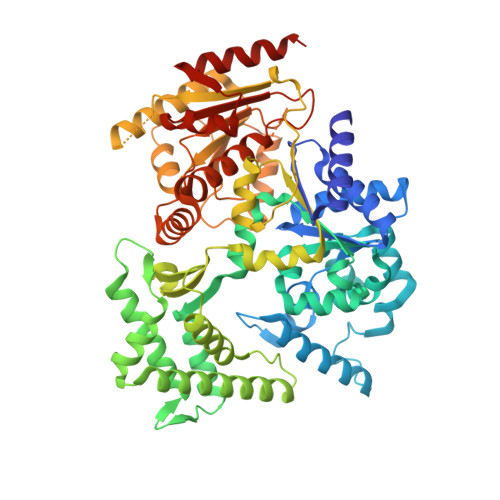
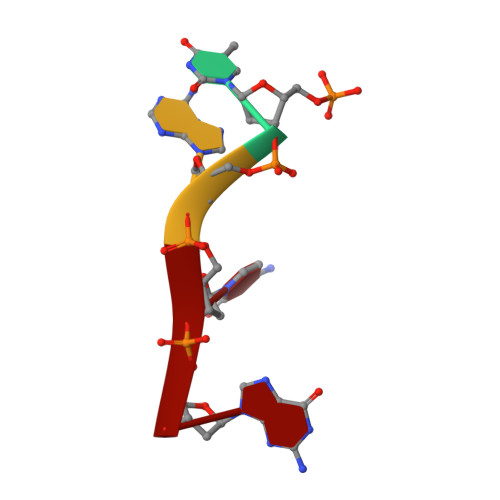
Functional and Structural Studies of the Nucleotide Excision Repair Helicase Xpd Suggest a Polarity for DNA Translocation.
Kuper, J., Wolski, S.C., Michels, G., Kisker, C.(2011) EMBO J 31: 494
- PubMed: 22081108 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/emboj.2011.374
- Primary Citation Related Structures:
4A15 - PubMed Abstract:
The XPD protein is a vital subunit of the general transcription factor TFIIH which is not only involved in transcription but is also an essential component of the eukaryotic nucleotide excision DNA repair (NER) pathway. XPD is a superfamily-2 5'-3' helicase containing an iron-sulphur cluster. Its helicase activity is indispensable for NER and it plays a role in the damage verification process. Here, we report the first structure of XPD from Thermoplasma acidophilum (taXPD) in complex with a short DNA fragment, thus revealing the polarity of the translocated strand and providing insights into how the enzyme achieves its 5'-3' directionality. Accompanied by a detailed mutational and biochemical analysis of taXPD, we define the path of the translocated DNA strand through the protein and identify amino acids that are critical for protein function.
- Rudolf Virchow Center for Experimental Biomedicine, Institute for Structural Biology, University of Würzburg, Würzburg, Germany. jochen.kuper@virchow.uni-wuerzburg.de
Organizational Affiliation: