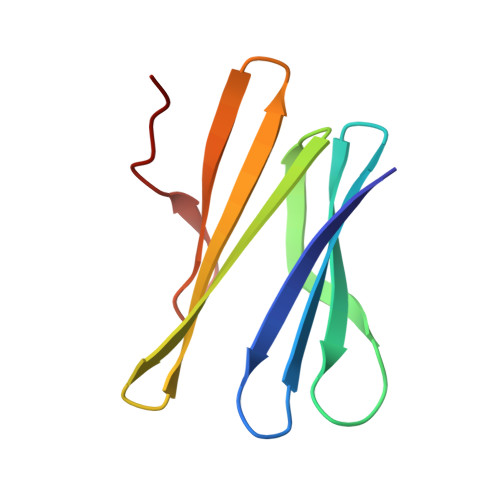
Fucose-Binding Lectin from Opportunistic Pathogen Burkholderia Ambifaria Binds to Both Plant and Human Oligosaccharidic Epitopes.
Audfray, A., Claudinon, J., Abounit, S., Ruvoen-Clouet, N., Larson, G., Smith, D.F., Wimmerova, M., Le Pendu, J., Romer, W., Varrot, A., Imberty, A.(2012) J Biological Chem 287: 4335
- PubMed: 22170069 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M111.314831
- Primary Citation Related Structures:
3ZW0, 3ZW2, 3ZWE, 3ZZV - PubMed Abstract:
Burkholderia ambifaria is generally associated with the rhizosphere of plants where it has biocontrol effects on other microorganisms. It is also a member of the Burkholderia cepacia complex, a group of closely related bacteria that cause lung infections in immunocompromised patients as well as in patients with granulomatous disease or cystic fibrosis. Our previous work indicated that fucose on human epithelia is a frequent target for lectins and adhesins of lung pathogens (Sulák, O., Cioci, G., Lameignère, E., Balloy, V., Round, A., Gutsche, I., Malinovská, L., Chignard, M., Kosma, P., Aubert, D. F., Marolda, C. L., Valvano, M. A., Wimmerová, M., and Imberty, A. (2011) PLoS Pathog. 7, e1002238). Analysis of the B. ambifaria genome identified BambL as a putative fucose-binding lectin. The 87-amino acid protein was produced recombinantly and demonstrated to bind to fucosylated oligosaccharides with a preference for αFuc1-2Gal epitopes. Crystal structures revealed that it associates as a trimer with two fucose-binding sites per monomer. The overall fold is a six-bladed β-propeller formed by oligomerization as in the Ralstonia solanacearum lectin and not by sequential domains like the fungal fucose lectin from Aleuria aurantia. The affinity of BambL for small fucosylated glycans is very high as demonstrated by microcalorimetry (K(D) < 1 μM). Plant cell wall oligosaccharides and human histo-blood group oligosaccharides H-type 2 and Lewis Y are bound with equivalent efficiency. Binding to artificial glycosphingolipid-containing vesicles, human saliva, and lung tissues confirmed that BambL could recognize a wide spectrum of fucosylated epitopes, albeit with a lower affinity for biological material from nonsecretor individuals.
- Centre de Recherche sur les Macromolécules Végétales (CERMAV)-CNRS, Université Joseph Fourier and Institut de Chimie Moléculaire de Grenoble, 38041 Grenoble, France.
Organizational Affiliation: