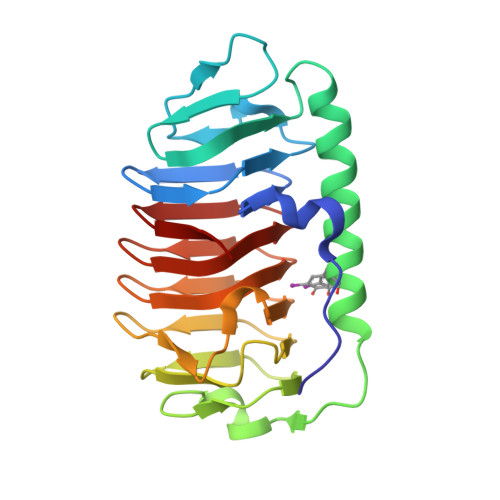
Ice-binding site of snow mold fungus antifreeze protein deviates from structural regularity and high conservation
Kondo, H., Hanada, Y., Sugimoto, H., Hoshino, T., Garnham, C.P., Davies, P.L., Tsuda, S.(2012) Proc Natl Acad Sci U S A 109: 9360-9365
- PubMed: 22645341 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1121607109
- Primary Citation Related Structures:
3VN3 - PubMed Abstract:
Antifreeze proteins (AFPs) are found in organisms ranging from fish to bacteria, where they serve different functions to facilitate survival of their host. AFPs that protect freeze-intolerant fish and insects from internal ice growth bind to ice using a regular array of well-conserved residues/motifs. Less is known about the role of AFPs in freeze-tolerant species, which might be to beneficially alter the structure of ice in or around the host. Here we report the 0.95-Å high-resolution crystal structure of a 223-residue secreted AFP from the snow mold fungus Typhula ishikariensis. Its main structural element is an irregular β-helix with six loops of 18 or more residues that lies alongside an α-helix. β-Helices have independently evolved as AFPs on several occasions and seem ideally structured to bind to several planes of ice, including the basal plane. A novelty of the β-helical fold is the nonsequential arrangement of loops that places the N- and C termini inside the solenoid of β-helical coils. The ice-binding site (IBS), which could not be predicted from sequence or structure, was located by site-directed mutagenesis to the flattest surface of the protein. It is remarkable for its lack of regularity and its poor conservation in homologs from psychrophilic diatoms and bacteria and other fungi.
- Bioproduction Research Institute, National Institute of Advanced Industrial Science and Technology, Sapporo 062-8517, Japan.
Organizational Affiliation: