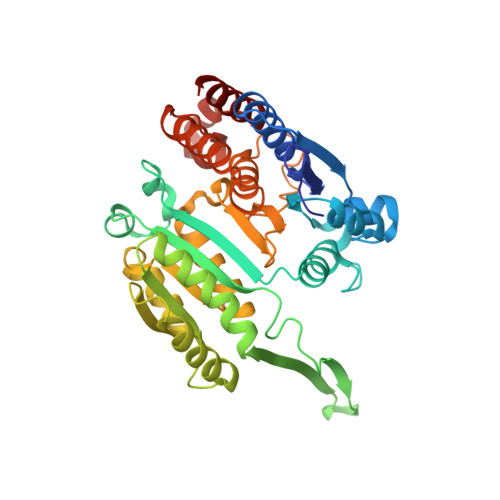
Structural analysis of 3-isopropylmalate dehydrogenase from the obligate piezophile Shewanella benthica DB21MT-2 and the nonpiezophile Shewanella oneidensis MR-1
Nagae, T., Kato, C., Watanabe, N.(2012) Acta Crystallogr Sect F Struct Biol Cryst Commun 68: 265-268
- PubMed: 22442218
- DOI: https://doi.org/10.1107/S1744309112001443
- Primary Citation Related Structures:
3VMJ, 3VMK - PubMed Abstract:
Organisms living in deep seas such as the Mariana Trench must be adapted to the extremely high pressure environment. For example, the 3-isopropylmalate dehydrogenase from the obligate piezophile Shewanella benthica DB21MT-2 (SbIPMDH) remains active in extreme conditions under which that from the land bacterium S. oneidensis MR-1 (SoIPMDH) becomes inactivated. In order to unravel the differences between these two IPMDHs, their structures were determined at ~1.5 Å resolution. Comparison of the structures of the two enzymes shows that SbIPMDH is in a more open form and has a larger internal cavity volume than SoIPMDH at atmospheric pressure. This loosely packed structure of SbIPMDH could help it to avoid pressure-induced distortion of the native structure and to remain active at higher pressures than SoIPMDH.
- Department of Biotechnology, Graduate School of Engineering, Nagoya University, Japan.
Organizational Affiliation: