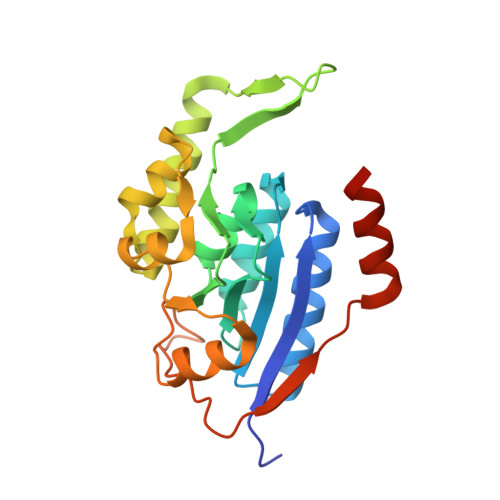
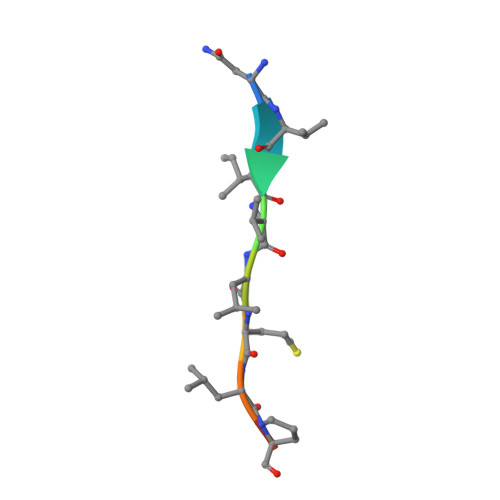
Crystal structure of a membrane stomatin-specific protease in complex with a substrate Peptide
Yokoyama, H., Takizawa, N., Kobayashi, D., Matsui, I., Fujii, S.(2012) Biochemistry 51: 3872-3880
- PubMed: 22475127
- DOI: https://doi.org/10.1021/bi300098k
- Primary Citation Related Structures:
3VIV - PubMed Abstract:
Membrane-bound proteases are involved in various regulatory functions. A previous report indicated that the N-terminal region of PH1510p (1510-N) from the hyperthermophilic archaeon Pyrococcus horikoshii is a serine protease with a catalytic Ser-Lys dyad (Ser97 and Lys138) and specifically cleaves the C-terminal hydrophobic region of the p-stomatin PH1511p. In humans, an absence of stomatin is associated with a form of hemolytic anemia known as hereditary stomatocytosis. Here, the crystal structure of 1510-N K138A in complex with a peptide substrate was determined at 2.25 Å resolution. In the structure, a 1510-N dimer binds to one peptide. The six central residues (VIVLML) of the peptide are hydrophobic and in a pseudopalindromic structure and therefore favorably fit into the hydrophobic active tunnel of the 1510-N dimer, although 1510-N degrades the substrate at only one point. A comparison with unliganded 1510-N K138A revealed that the binding of the substrate causes a large rotational and translational displacement between protomers and produces a tunnel suitable for binding the peptide. When the peptide binds, the flexible L2 loop of one protomer forms β-strands, whereas that of the other protomer remains in a loop form, indicating that one protomer binds to the peptide more tightly than the other protomer. The Ala138 residues of the two protomers are located very close together (the distance between the two Cβ atoms is 3.6 Å). Thus, in wild-type 1510-N, the close positioning of the catalytic Ser97 and Lys138 residues may be induced by electrostatic repulsion of the two Lys138 side chains of the protomers.
- School of Pharmaceutical Sciences, University of Shizuoka, Shizuoka 422-8526, Japan. h-yokoya@u-shizuoka-ken.ac.jp
Organizational Affiliation: