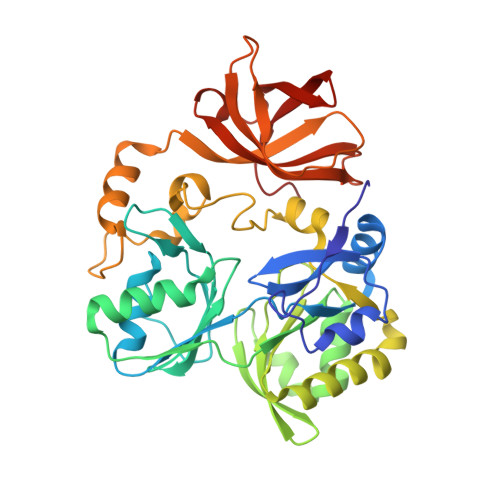
Structures of dimethylsulfoniopropionate-dependent demethylase from the marine organism Pelagabacter ubique.
Schuller, D.J., Reisch, C.R., Moran, M.A., Whitman, W.B., Lanzilotta, W.N.(2012) Protein Sci 21: 289-298
- PubMed: 22162093 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.2015
- Primary Citation Related Structures:
3TFH, 3TFI, 3TFJ - PubMed Abstract:
Dimethylsulfoniopropionate (DMSP) is a ubiquitous algal metabolite and common carbon and sulfur source for marine bacteria. DMSP is a precursor for the climatically active gas dimethylsulfide that is readily oxidized to sulfate, sulfur dioxide, methanesulfonic acid, and other products that act as cloud condensation nuclei. Although the environmental importance of DMSP metabolism has been known for some time, the enzyme responsible for DMSP demethylation by marine bacterioplankton, dimethylsufoniopropionate-dependent demethylase A (DmdA, EC 2.1.1.B5), has only recently been identified and biochemically characterized. In this work, we report the structure for the apoenzyme DmdA from Pelagibacter ubique (2.1 Å), as well as for DmdA co-crystals soaked with substrate DMSP (1.6 Å) or the cofactor tetrahydrofolate (THF) (1.6 Å). Surprisingly, the overall fold of the DmdA is not similar to other enzymes that typically utilize the reduced form of THF and in fact is a triple domain structure similar to what has been observed for the glycine cleavage T protein or sarcosine oxidase. Specifically, while the THF binding fold appears conserved, previous biochemical studies have shown that all enzymes with a similar fold produce 5,10-methylene-THF, while DmdA catalyzes a redox-neutral methyl transfer reaction to produce 5-methyl-THF. On the basis of the findings presented herein and the available biochemical data, we outline a mechanism for a redox-neutral methyl transfer reaction that is novel to this conserved THF binding domain.
- Cornell High Energy Synchrotron Source, Cornell University, Ithaca, New York 14853, USA.
Organizational Affiliation: