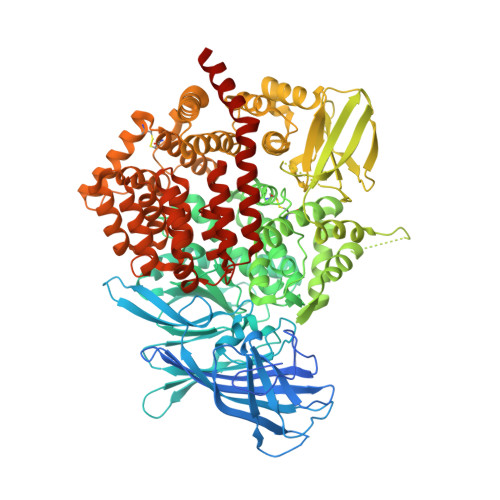
The crystal structure of human endoplasmic reticulum aminopeptidase 2 reveals the atomic basis for distinct roles in antigen processing.
Birtley, J.R., Saridakis, E., Stratikos, E., Mavridis, I.M.(2012) Biochemistry 51: 286-295
- PubMed: 22106953 Search on PubMed
- DOI: https://doi.org/10.1021/bi201230p
- Primary Citation Related Structures:
3SE6 - PubMed Abstract:
Endoplasmic reticulum aminopeptidases ERAP1 and ERAP2 cooperate to trim a vast variety of antigenic peptide precursors to generate mature epitopes for binding to major histocompatibility class I molecules. We report here the first structure of ERAP2 determined at 3.08 Å by X-ray crystallography. On the basis of residual electron density, a lysine residue has been modeled in the active site of the enzyme; thus, the structure corresponds to an enzyme-product complex. The overall domain organization is highly similar to that of the recently determined structure of ERAP1 in its closed conformation. A large internal cavity adjacent to the catalytic site can accommodate large peptide substrates. The ERAP2 structure provides a structural explanation for the different peptide N-terminal specificities between ERAP1 and ERAP2 and suggests that such differences extend throughout the whole peptide sequence. A noncrystallographic dimer observed may constitute a model for a proposed ERAP1-ERAP2 heterodimer. Overall, the structure helps explain how two homologous aminopeptidases cooperate to process a large variety of sequences, a key property of their biological role.
- Structural and Supramolecular Chemistry Laboratory, Institute of Physical Chemistry, National Center for Scientific Research Demokritos, Aghia Paraskevi 15310, Athens, Greece.
Organizational Affiliation: