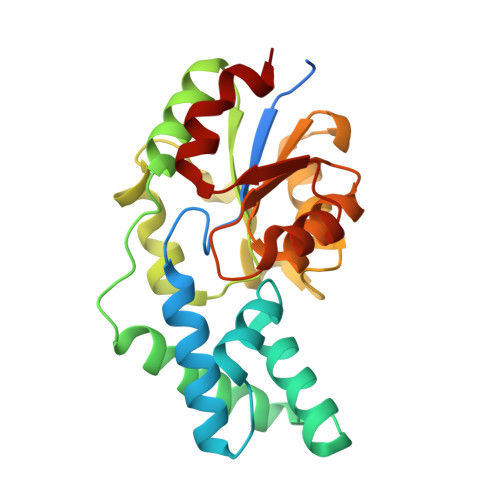
Divergence of Structure and Function in the Haloacid Dehalogenase Enzyme Superfamily: Bacteroides thetaiotaomicron BT2127 Is an Inorganic Pyrophosphatase.
Huang, H., Patskovsky, Y., Toro, R., Farelli, J.D., Pandya, C., Almo, S.C., Allen, K.N., Dunaway-Mariano, D.(2011) Biochemistry 50: 8937-8949
- PubMed: 21894910
- DOI: https://doi.org/10.1021/bi201181q
- Primary Citation of Related Structures:
3QU2, 3QU4, 3QU5, 3QU7, 3QU9, 3QUB, 3QUC, 3QUQ, 3QUT, 3QX7, 3QXG, 3QYP, 3R9K - PubMed Abstract:
The explosion of protein sequence information requires that current strategies for function assignment evolve to complement experimental approaches with computationally based function prediction. This necessitates the development of strategies based on the identification of sequence markers in the form of specificity determinants and a more informed definition of orthologues. Herein, we have undertaken the function assignment of the unknown haloalkanoate dehalogenase superfamily member BT2127 (Uniprot accession code Q8A5 V9) from Bacteroides thetaiotaomicron using an integrated bioinformatics-structure-mechanism approach. The substrate specificity profile and steady-state rate constants of BT2127 (with a k(cat)/K(m) value for pyrophosphate of ~1 × 10(5) M(-1) s(-1)), together with the gene context, support the assigned in vivo function as an inorganic pyrophosphatase. The X-ray structural analysis of wild-type BT2127 and several variants generated by site-directed mutagenesis shows that substrate discrimination is based, in part, on active site space restrictions imposed by the cap domain (specifically by residues Tyr76 and Glu47). Structure-guided site-directed mutagenesis coupled with kinetic analysis of the mutant enzymes identified the residues required for catalysis, substrate binding, and domain-domain association. On the basis of this structure-function analysis, the catalytic residues Asp11, Asp13, Thr113, and Lys147 as well the metal binding residues Asp171, Asn172, and Glu47 were used as markers to confirm BT2127 orthologues identified via sequence searches. This bioinformatic analysis demonstrated that the biological range of BT2127 orthologue is restricted to the phylum Bacteroidetes/Chlorobi. The key structural determinants in the divergence of BT2127 and its closest homologue, β-phosphoglucomutase, control the leaving group size (phosphate vs glucose phosphate) and the position of the Asp acid/base in the open versus closed conformations. HADSF pyrophosphatases represent a third mechanistic and fold type for bacterial pyrophosphatases.
- Department of Chemistry and Chemical Biology, University of New Mexico, Albuquerque, New Mexico 87131, USA.
Organizational Affiliation: