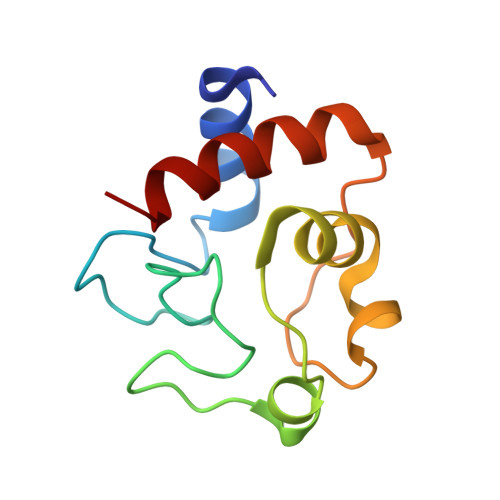
Nitrate as a probe of cytochrome c surface: crystallographic identification of crucial "hot spots" for protein-protein recognition.
De March, M., Demitri, N., De Zorzi, R., Casini, A., Gabbiani, C., Guerri, A., Messori, L., Geremia, S.(2014) J Inorg Biochem 135: 58-67
- PubMed: 24662464 Search on PubMed
- DOI: https://doi.org/10.1016/j.jinorgbio.2014.02.015
- Primary Citation Related Structures:
3O1Y, 3O20 - PubMed Abstract:
The electrostatic surface of cytochrome c and its changes with the iron oxidation state are involved in the docking and undocking processes of this protein to its biological partners in the mitochondrial respiratory pathway. To investigate the subtle mechanisms of formation of productive macromolecular complexes and of their breakage following the electron transfer process, the X-ray structures of horse heart ferri-cytochrome c (trigonal form) and ferro-cytochrome c (monoclinic form) were obtained using nitrate ions both as a crystallizing agent and an anionic probe for mapping the electrostatic surface changes. Both crystal forms contain three protein molecules in the asymmetric unit. In addition, a total of 21.5 and 18 crystallographically independent nitrate ions were identified for the trigonal and monoclinic forms, respectively. By matching all the six crystallographically independent protein molecules, 26 different anion-protein interaction sites were identified on the surfaces of cytochrome c, 10 of which were found in both forms, 8 present only in the oxidized and 8 only in the reduced form. The structural analysis of the electron transfer complexes, based on this new information, suggests a specific exit strategy for cytochrome c after formation of productive protein-protein complexes: a directional sliding mechanism for the electron shuttle on the surface of the redox partner is proposed to take place after the electron transfer process has occurred.
- Centre of Excellence in Biocrystallography, Department of Chemical and Pharmaceutical Sciences, University of Trieste, via L. Giorgeri 1, 34127 Trieste, Italy.
Organizational Affiliation: