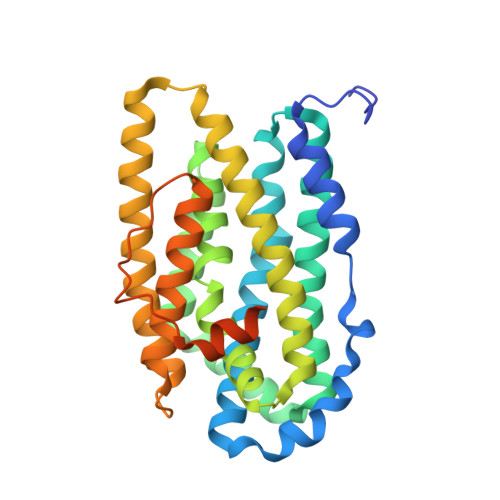
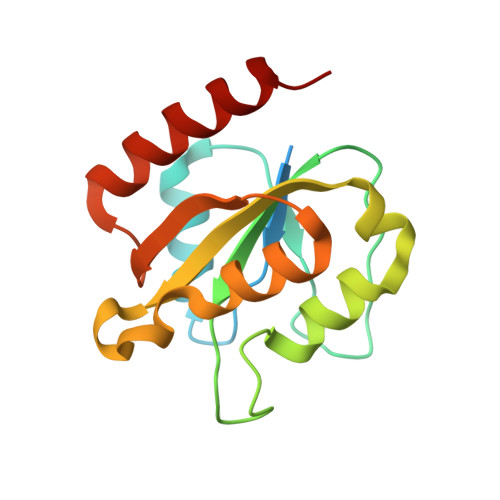
Structural basis for activation of class Ib ribonucleotide reductase.
Boal, A.K., Cotruvo, J.A., Stubbe, J., Rosenzweig, A.C.(2010) Science 329: 1526-1530
- PubMed: 20688982 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/science.1190187
- Primary Citation Related Structures:
3N37, 3N38, 3N39, 3N3A, 3N3B - PubMed Abstract:
The class Ib ribonucleotide reductase of Escherichia coli can initiate reduction of nucleotides to deoxynucleotides with either a Mn(III)2-tyrosyl radical (Y•) or a Fe(III)2-Y• cofactor in the NrdF subunit. Whereas Fe(III)2-Y• can self-assemble from Fe(II)2-NrdF and O2, activation of Mn(II)2-NrdF requires a reduced flavoprotein, NrdI, proposed to form the oxidant for cofactor assembly by reduction of O2. The crystal structures reported here of E. coli Mn(II)2-NrdF and Fe(II)2-NrdF reveal different coordination environments, suggesting distinct initial binding sites for the oxidants during cofactor activation. In the structures of Mn(II)2-NrdF in complex with reduced and oxidized NrdI, a continuous channel connects the NrdI flavin cofactor to the NrdF Mn(II)2 active site. Crystallographic detection of a putative peroxide in this channel supports the proposed mechanism of Mn(III)2-Y• cofactor assembly.
- Department of Biochemistry, Molecular Biology and Cell Biology, Northwestern University, Evanston, IL 60208, USA.
Organizational Affiliation: