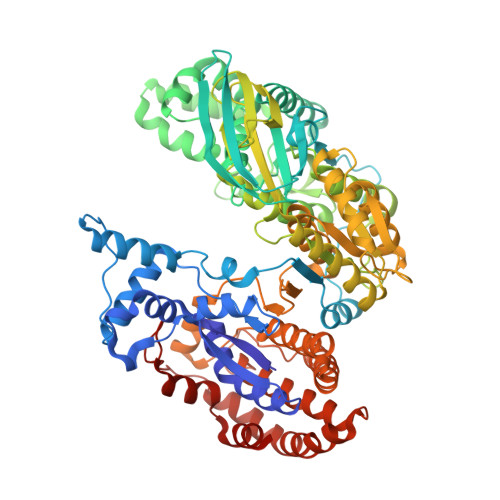
Structure of a highly NADP+-specific isocitrate dehydrogenase.
Sidhu, N.S., Delbaere, L.T., Sheldrick, G.M.(2011) Acta Crystallogr D Biol Crystallogr 67: 856-869
- PubMed: 21931217 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444911028575
- Primary Citation Related Structures:
3MBC - PubMed Abstract:
Isocitrate dehydrogenase catalyzes the first oxidative and decarboxylation steps in the citric acid cycle. It also lies at a crucial bifurcation point between CO2-generating steps in the cycle and carbon-conserving steps in the glyoxylate bypass. Hence, the enzyme is a focus of regulation. The bacterial enzyme is typically dependent on the coenzyme nicotinamide adenine dinucleotide phosphate. The monomeric enzyme from Corynebacterium glutamicum is highly specific towards this coenzyme and the substrate isocitrate while retaining a high overall efficiency. Here, a 1.9 Å resolution crystal structure of the enzyme in complex with its coenzyme and the cofactor Mg2+ is reported. Coenzyme specificity is mediated by interactions with the negatively charged 2'-phosphate group, which is surrounded by the side chains of two arginines, one histidine and, via a water, one lysine residue, forming ion pairs and hydrogen bonds. Comparison with a previous apoenzyme structure indicates that the binding site is essentially preconfigured for coenzyme binding. In a second enzyme molecule in the asymmetric unit negatively charged aspartate and glutamate residues from a symmetry-related enzyme molecule interact with the positively charged arginines, abolishing coenzyme binding. The holoenzyme from C. glutamicum displays a 36° interdomain hinge-opening movement relative to the only previous holoenzyme structure of the monomeric enzyme: that from Azotobacter vinelandii. As a result, the active site is not blocked by the bound coenzyme as in the closed conformation of the latter, but is accessible to the substrate isocitrate. However, the substrate-binding site is disrupted in the open conformation. Hinge points could be pinpointed for the two molecules in the same crystal, which show a 13° hinge-bending movement relative to each other. One of the two pairs of hinge residues is intimately flanked on both sides by the isocitrate-binding site. This suggests that binding of a relatively small substrate (or its competitive inhibitors) in tight proximity to a hinge point could lead to large conformational changes leading to a closed, presumably catalytically active (or inactive), conformation. It is possible that the small-molecule concerted inhibitors glyoxylate and oxaloacetate similarly bind close to the hinge, leading to an inactive conformation of the enzyme.
- Department of Structural Chemistry, University of Göttingen, Tammannstrasse 4, D-37077 Göttingen, Germany. nsidhu@shelx.uni-ac.gwdg.de
Organizational Affiliation: