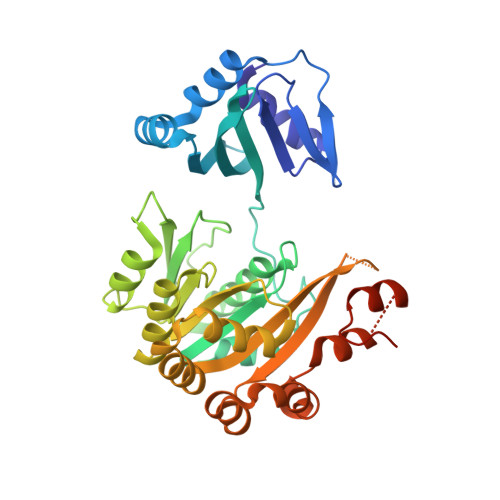
P. aeruginosa PilT structures with and without nucleotide reveal a dynamic type IV pilus retraction motor.
Misic, A.M., Satyshur, K.A., Forest, K.T.(2010) J Mol Biology 400: 1011-1021
- PubMed: 20595000 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2010.05.066
- Primary Citation Related Structures:
3JVU, 3JVV - PubMed Abstract:
Type IV pili are bacterial extracellular filaments that can be retracted to create force and motility. Retraction is accomplished by the motor protein PilT. Crystal structures of Pseudomonas aeruginosa PilT with and without bound beta,gamma-methyleneadenosine-5'-triphosphate have been solved at 2.6 A and 3.1 A resolution, respectively, revealing an interlocking hexamer formed by the action of a crystallographic 2-fold symmetry operator on three subunits in the asymmetric unit and held together by extensive ionic interactions. The roles of two invariant carboxylates, Asp Box motif Glu163 and Walker B motif Glu204, have been assigned to Mg(2+) binding and catalysis, respectively. The nucleotide ligands in each of the subunits in the asymmetric unit of the beta,gamma-methyleneadenosine-5'-triphosphate-bound PilT are not equally well ordered. Similarly, the three subunits in the asymmetric unit of both structures exhibit differing relative conformations of the two domains. The 12 degrees and 20 degrees domain rotations indicate motions that occur during the ATP-coupled mechanism of the disassembly of pili into membrane-localized pilin monomers. Integrating these observations, we propose a three-state "Ready, Active, Release" model for the action of PilT.
- Department of Biomolecular Chemistry, University of Wisconsin-Madison, Madison, 1550 Linden Drive, Madison, WI 53706, USA.
Organizational Affiliation: