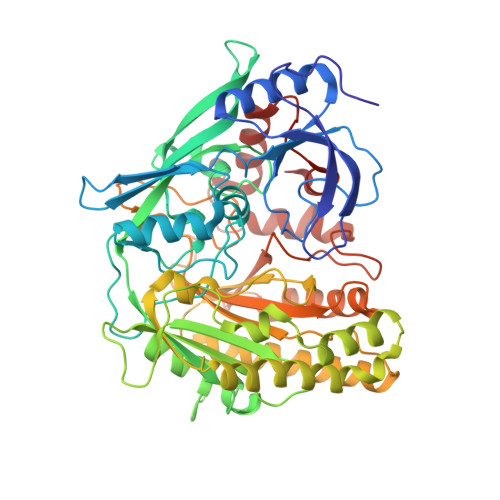
Structural characterization of the organic solvent-stable cholesterol oxidase from Chromobacterium sp. DS-1.
Sagermann, M., Ohtaki, A., Newton, K., Doukyu, N.(2010) J Struct Biol 170: 32-40
- PubMed: 20102741 Search on PubMed
- DOI: https://doi.org/10.1016/j.jsb.2010.01.012
- Primary Citation Related Structures:
3JS8 - PubMed Abstract:
Cholesterol oxidase is of significant commercial interest as it is widely used as a biosensor for the detection of cholesterol in clinical samples, blood serum and food. Increased stability of this enzyme with regards to temperature and different solvent conditions are of great importance to the reliability and versatility of its applications. We here report the crystal structure of the cholesterol oxidase of Chromobacterium sp. DS-1 (CHOLOX). In contrast to other previously characterized cholesterol oxidases, this enzyme retains high activity in organic solvents and detergents at temperatures above 85 degrees C despite its mesophilic origin. With the availability of one other homologous oxidase of known three-dimensional structure, a detailed comparison of its sequence and structure was performed to elucidate the mechanisms of stabilization. In contrast to factors that typically contribute to the stability of thermophilic proteins, the structure of CHOLOX exhibits a larger overall cavity volume, less charged residues and less salt bridge interactions. Moreover, the vast majority of residue substitutions were found on or near the protein's solvent exposed surface. We propose that the engineering of enhanced stability may also be accomplished through selective engineering of the protein periphery rather than by redesigning its entire core.
- Department of Chemistry and Biochemistry, University of California, Santa Barbara, CA 93106-9510, USA. Sagermann@chem.ucsb.edu
Organizational Affiliation: