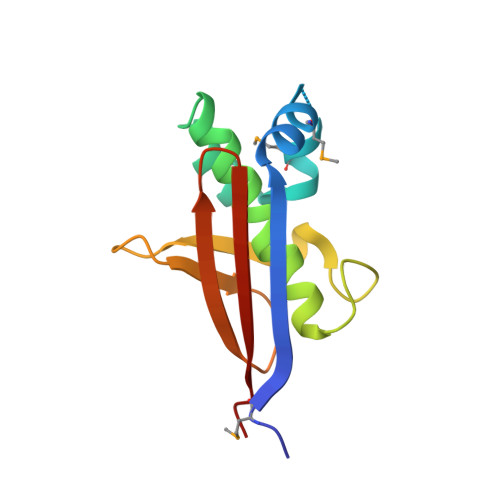
Structural characterization and comparison of three acyl-carrier-protein synthases from pathogenic bacteria.
Halavaty, A.S., Kim, Y., Minasov, G., Shuvalova, L., Dubrovska, I., Winsor, J., Zhou, M., Onopriyenko, O., Skarina, T., Papazisi, L., Kwon, K., Peterson, S.N., Joachimiak, A., Savchenko, A., Anderson, W.F.(2012) Acta Crystallogr D Biol Crystallogr 68: 1359-1370
- PubMed: 22993090 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S0907444912029101
- Primary Citation Related Structures:
3HYK, 3QMN, 4JM7 - PubMed Abstract:
Some bacterial type II fatty-acid synthesis (FAS II) enzymes have been shown to be important candidates for drug discovery. The scientific and medical quest for new FAS II protein targets continues to stimulate research in this field. One of the possible additional candidates is the acyl-carrier-protein synthase (AcpS) enzyme. Its holo form post-translationally modifies the apo form of an acyl carrier protein (ACP), which assures the constant delivery of thioester intermediates to the discrete enzymes of FAS II. At the Center for Structural Genomics of Infectious Diseases (CSGID), AcpSs from Staphylococcus aureus (AcpS(SA)), Vibrio cholerae (AcpS(VC)) and Bacillus anthracis (AcpS(BA)) have been structurally characterized in their apo, holo and product-bound forms, respectively. The structure of AcpS(BA) is emphasized because of the two 3',5'-adenosine diphosphate (3',5'-ADP) product molecules that are found in each of the three coenzyme A (CoA) binding sites of the trimeric protein. One 3',5'-ADP is bound as the 3',5'-ADP part of CoA in the known structures of the CoA-AcpS and 3',5'-ADP-AcpS binary complexes. The position of the second 3',5'-ADP has never been described before. It is in close proximity to the first 3',5'-ADP and the ACP-binding site. The coordination of two ADPs in AcpS(BA) may possibly be exploited for the design of AcpS inhibitors that can block binding of both CoA and ACP.
- Center for Structural Genomics of Infectious Diseases, USA.
Organizational Affiliation: