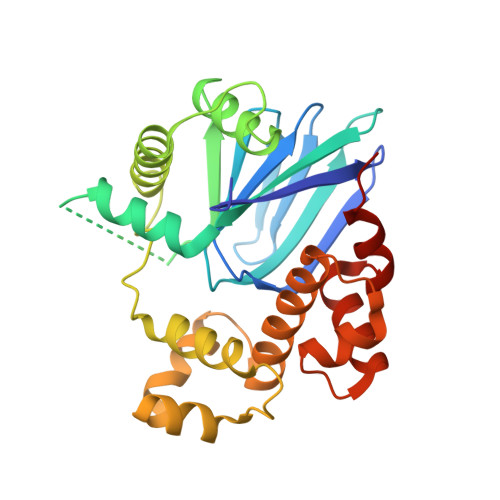
Structure of Vibrio cholerae ToxT reveals a mechanism for fatty acid regulation of virulence genes.
Lowden, M.J., Skorupski, K., Pellegrini, M., Chiorazzo, M.G., Taylor, R.K., Kull, F.J.(2010) Proc Natl Acad Sci U S A 107: 2860-2865
- PubMed: 20133655 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0915021107
- Primary Citation Related Structures:
3GBG - PubMed Abstract:
Cholera is an acute intestinal infection caused by the bacterium Vibrio cholerae. In order for V. cholerae to cause disease, it must produce two virulence factors, the toxin-coregulated pilus (TCP) and cholera toxin (CT), whose expression is controlled by a transcriptional cascade culminating with the expression of the AraC-family regulator, ToxT. We have solved the 1.9 A resolution crystal structure of ToxT, which reveals folds in the N- and C-terminal domains that share a number of features in common with AraC, MarA, and Rob as well as the unexpected presence of a buried 16-carbon fatty acid, cis-palmitoleate. The finding that cis-palmitoleic acid reduces TCP and CT expression in V. cholerae and prevents ToxT from binding to DNA in vitro provides a direct link between the host environment of V. cholerae and regulation of virulence gene expression.
- Department of Chemistry, Dartmouth College, 6128 Burke Laboratory, Hanover, NH 03755, USA.
Organizational Affiliation: