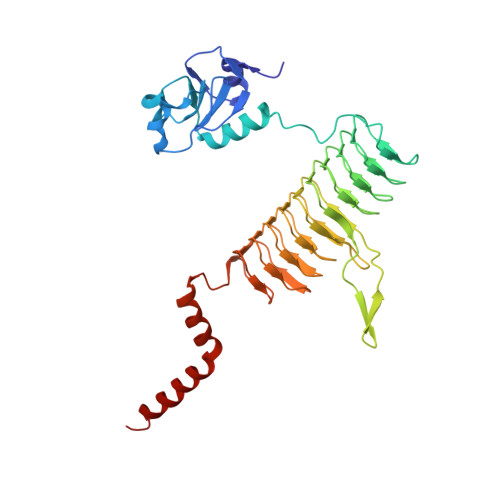
Crystal structure and acyl chain selectivity of Escherichia coli LpxD, the N-acyltransferase of lipid A biosynthesis
Bartling, C.M., Raetz, C.R.(2009) Biochemistry 48: 8672-8683
- PubMed: 19655786 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/bi901025v
- Primary Citation Related Structures:
3EH0 - PubMed Abstract:
LpxD catalyzes the third step of lipid A biosynthesis, the R-3-hydroxyacyl-ACP-dependent N-acylation of UDP-3-O-(acyl)-alpha-D-glucosamine, and is a target for new antibiotic development. Here we report the 2.6 A crystal structure of the Escherichia coli LpxD homotrimer (EcLpxD). As is the case in Chlamydia trachomatis LpxD (CtLxpD), each EcLpxD chain consists of an N-terminal uridine-binding region, a left-handed parallel beta-helix (LbetaH), and a C-terminal alpha-helical domain. The backbones of the LbetaH domains of the two enzymes are similar, as are the positions of key active site residues. The N-terminal nucleotide binding domains are oriented differently relative to the LbetaH regions, but are similar when overlaid on each other. The orientation of the EcLpxD tripeptide (residues 303-305), connecting the distal end of the LbetaH and the proximal end of the C-terminal helical domains, differs from its counterpart in CtLpxD (residues 311-312); this results in a 120 degrees rotation of the C-terminal domain relative to the LbetaH region in EcLpxD versus CtLpxD. M290 of EcLpxD appears to cap the distal end of a hydrophobic cleft that binds the acyl chain of the R-3-hydroxyacyl-ACP donor substrate. Under standard assay conditions, wild-type EcLpxD prefers R,S-3-hydroxymyristoyl-ACP over R,S-3-hydroxypalmitoyl-ACP by a factor of 3, whereas the M290A mutant has the opposite selectivity. Both wild-type and M290A EcLpxD rescue the conditional lethality of E. coli RL25, a temperature-sensitive strain harboring point mutations in lpxD. Complementation with wild-type EcLpxD restores normal lipid A containing only N-linked hydroxymyristate to RL25 at 42 degrees C, as judged by mass spectrometry, whereas the M290A mutant generates multiple lipid A species containing one or two longer hydroxy fatty acids in place of the usual R-3-hydroxymyristate at positions 2 and 2'.
- Department of Biochemistry, Duke University Medical Center, Durham, North Carolina 27710, USA.
Organizational Affiliation: