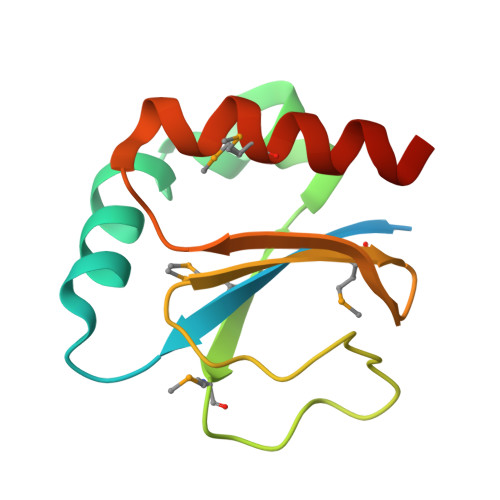
The structure of the periplasmic thiol-disulfide oxidoreductase SoxS from Paracoccus pantotrophus indicates a triple Trx/Grx/DsbC functionality in chemotrophic sulfur oxidation.
Carius, Y., Rother, D., Friedrich, C.G., Scheidig, A.J.(2009) Acta Crystallogr D Biol Crystallogr 65: 229-240
- PubMed: 19237745 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444908043023
- Primary Citation Related Structures:
3D4T, 3DML - PubMed Abstract:
The periplasmic thiol-disulfide oxidoreductase SoxS is beneficial for the sulfur-oxidizing (Sox) phenotype of the facultative chemotrophic bacterium Paracoccus pantotrophus and is not part of the Sox enzyme system. SoxS combines features of thioredoxins, glutaredoxins and the thiol-disulfide oxidoreductases of the Dsb family in structure, target specificity and reaction. The structure of SoxS was solved in oxidized and reduced forms at 2.1 and 1.9 A resolution, respectively. SoxS revealed high structural homology to typical cytoplasmic bacterial thioredoxins. In contrast, SoxS contained the active-site motif Pro-Gly-Cys-Leu-Tyr-Cys that is not present in other thioredoxins. Interestingly, the sequence of this motif is closely related to the Pro-Gly-Cys-Pro-Tyr-Cys sequence of some glutaredoxins and to the Pro-Xaa-Cys-Xaa-Tyr-Cys sequences of some members of the DsbC and DsbG subfamilies of thiol-disulfide oxidoreductases. Furthermore, the proposed substrate of SoxS, the interprotein disulfide of SoxY, Cys110(Y)-Cys110(Y), is structurally similar to oxidized glutathione. However, SoxS is proposed to specifically reduce the interprotein disulfide between two SoxY subunits, releasing a heterodimeric SoxYZ as an active part of the sulfur-oxidation cycle.
- Abteilung für Strukturbiologie, Zoologisches Institut, Christian-Albrechts-Universität zu Kiel, Kiel, Germany.
Organizational Affiliation: