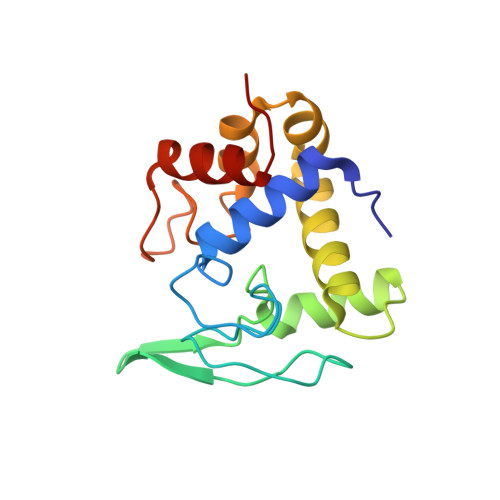
Crystal structure of the lytic transglycosylase from bacteriophage lambda in complex with hexa-N-acetylchitohexaose
Leung, A.K.W., Duewel, H.S., Honek, J.F., Berghuis, A.M.(2001) Biochemistry 40: 5665-5673
- PubMed: 11341831
- DOI: https://doi.org/10.1021/bi0028035
- Primary Citation Related Structures:
3D3D - PubMed Abstract:
The three-dimensional structure of the lytic transglycosylase from bacteriophage lambda, also known as bacteriophage lambda lysozyme, complexed to the hexasaccharide inhibitor, hexa-N-acetylchitohexaose, has been determined by X-ray crystallography at 2.6 A resolution. The unit cell contains two molecules of the lytic transglycosylase with two hexasaccharides bound. Each enzyme molecule is found to interact with four N-acetylglucosamine units from one hexasaccharide (subsites A-D) and two N-acetylglucosamine units from the second hexasaccharide (subsites E and F), resulting in all six subsites of the active site of this enzyme being filled. This crystallographic structure, therefore, represents the first example of a lysozyme in which all subsites are occupied, and detailed protein-oligosaccharide interactions are now available for this bacteriophage lytic transglycosylase. Examination of the active site furthermore reveals that of the two residues that have been implicated in the reaction mechanism of most other c-type lysozymes (Glu35 and Asp52 in hen egg white lysozyme), only a homologous Glu residue is present. The lambda lytic transglycosylase is therefore functionally closely related to the Escherichia coli Slt70 and Slt35 lytic transglycosylases and goose egg white lysozyme which also lack the catalytic aspartic acid.
- Antimicrobial Research Centre and Department of Biochemistry, McMaster University, Hamilton, Ontario L8N 3Z5, Canada.
Organizational Affiliation: