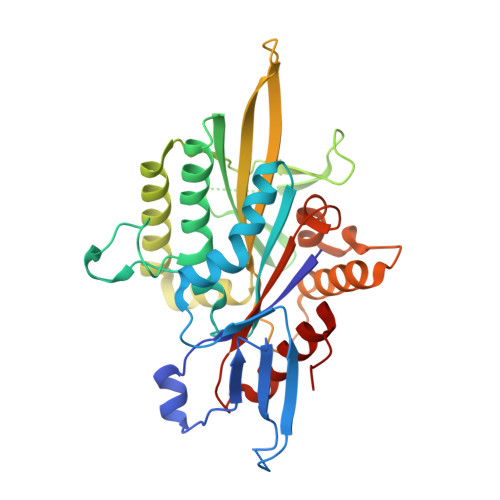
Structural Insights Into a Unique Inhibitor Binding Pocket in Kinesin Spindle Protein.
Ulaganathan, V., Talapatra, S.K., Rath, O., Pannifer, A.D., Hackney, D.D., Kozielski, F.(2013) J Am Chem Soc 135: 2263
- PubMed: 23305346 Search on PubMed
- DOI: https://doi.org/10.1021/ja310377d
- Primary Citation Related Structures:
3ZCW - PubMed Abstract:
Human kinesin Eg5 is a target for drug development in cancer chemotherapy with compounds in phase II clinical trials. These agents bind to a well-characterized allosteric pocket involving the loop L5 region, a structural element in kinesin-5 family members thought to provide inhibitor specificity. Using X-ray crystallography, kinetic, and biophysical methods, we have identified and characterized a distinct allosteric pocket in Eg5 able to bind inhibitors with nanomolar K(d). This pocket is formed by key structural elements thought to be pivotal for force generation in kinesins and may represent a novel site for therapeutic intervention in this increasingly well-validated drug target.
- The Molecular Motors Laboratory, The Beatson Institute for Cancer Research, Garscube Estate, Switchback Road, Glasgow G61 1BD, Scotland, UK.
Organizational Affiliation: