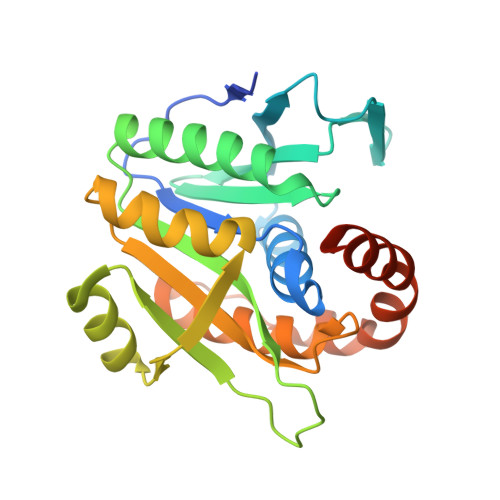
Crystal structure of archaeal homolog of proteasome-assembly chaperone PbaA
Sikdar, A., Satoh, T., Kawasaki, M., Kato, K.(2014) Biochem Biophys Res Commun 453: 493-497
- PubMed: 25285636 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2014.09.114
- Primary Citation Related Structures:
3WZ2 - PubMed Abstract:
Formation of the eukaryotic proteasome is not a spontaneous process but a highly ordered process assisted by several assembly chaperones. In contrast, archaeal proteasome subunits can spontaneously assemble into an active form. Recent bioinformatic analysis identified the proteasome-assembly chaperone-like proteins, PbaA and PbaB, in archaea. Our previous study showed that the PbaB homotetramer functions as a proteasome activator through its tentacle-like C-terminal segments. However, a functional role of the other homolog PbaA has remained elusive. Here we determined the 2.25-Å resolution structure of PbaA, illustrating its disparate tertiary and quaternary structures compared with PbaB. PbaA forms a homopentamer in which the C-terminal segments, with a putative proteasome-activating motif, are packed against the core. These findings offer deeper insights into the molecular evolution relationships between the proteasome-assembly chaperones and the proteasome activators.
- School of Physical Science, The Graduate University for Advanced Studies, 5-1 Higashiyama, Myodaiji, Okazaki, Aichi 444-8787, Japan; Okazaki Institute for Integrative Bioscience and Institute for Molecular Science, National Institutes of Natural Sciences, 5-1 Higashiyama, Myodaiji, Okazaki, Aichi 444-8787, Japan; Graduate School of Pharmaceutical Sciences, Nagoya City University, 3-1 Tanabe-dori, Mizuho-ku, Nagoya 467-8603, Japan.
Organizational Affiliation: